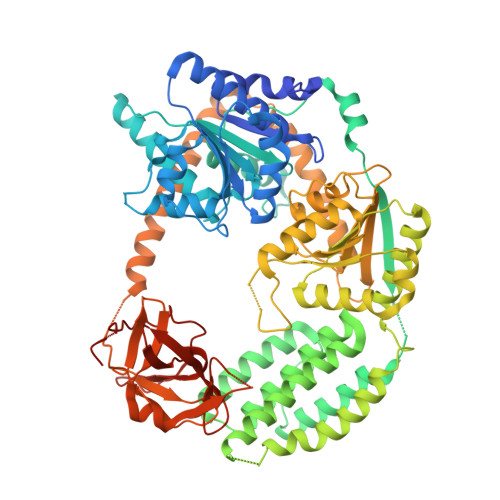
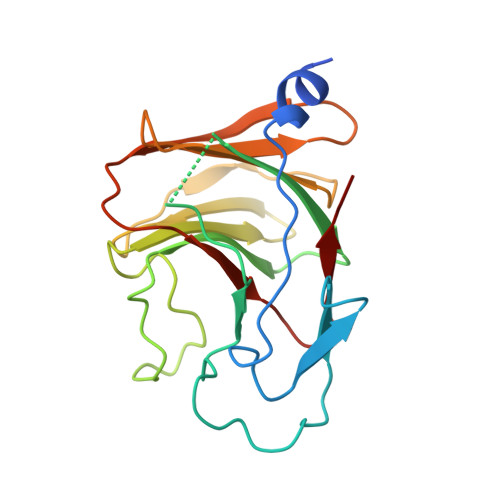
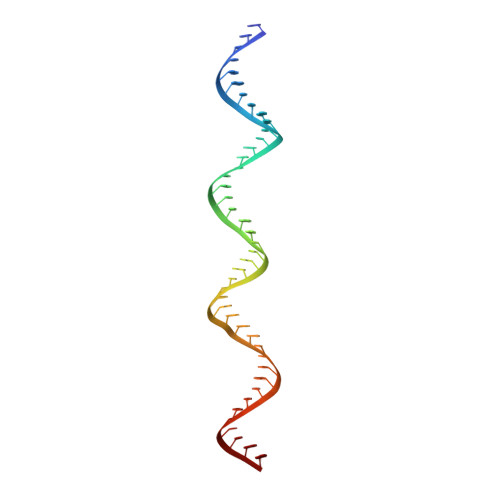Structural analysis of RIG-I-like receptors reveals ancient rules of engagement between diverse RNA helicases and TRIM ubiquitin ligases.
Kato, K., Ahmad, S., Zhu, Z., Young, J.M., Mu, X., Park, S., Malik, H.S., Hur, S.(2021) Mol Cell 81: 599-613.e8
- PubMed: 33373584 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2020.11.047
- Primary Citation Related Structures:
7JL0, 7JL1, 7JL2, 7JL3, 7JL4 - PubMed Abstract:
RNA helicases and E3 ubiquitin ligases mediate many critical functions in cells, but their actions have largely been studied in distinct biological contexts. Here, we uncover evolutionarily conserved rules of engagement between RNA helicases and tripartite motif (TRIM) E3 ligases that lead to their functional coordination in vertebrate innate immunity. Using cryoelectron microscopy and biochemistry, we show that RIG-I-like receptors (RLRs), viral RNA receptors with helicase domains, interact with their cognate TRIM/TRIM-like E3 ligases through similar epitopes in the helicase domains. Their interactions are avidity driven, restricting the actions of TRIM/TRIM-like proteins and consequent immune activation to RLR multimers. Mass spectrometry and phylogeny-guided biochemical analyses further reveal that similar rules of engagement may apply to diverse RNA helicases and TRIM/TRIM-like proteins. Our analyses suggest not only conserved substrates for TRIM proteins but also, unexpectedly, deep evolutionary connections between TRIM proteins and RNA helicases, linking ubiquitin and RNA biology throughout animal evolution.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA; Program in Cellular and Molecular Medicine, Boston Children's Hospital, Boston, MA 02115, USA.
Organizational Affiliation: