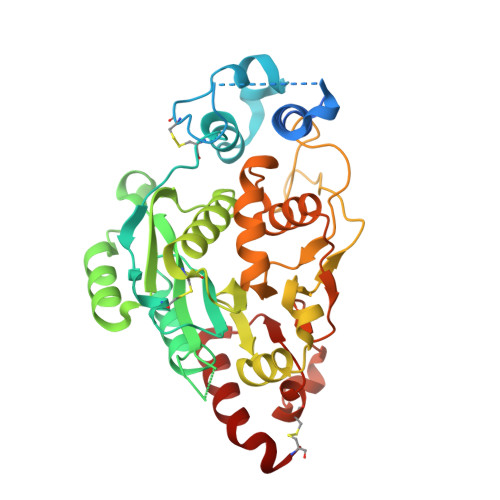
Structures and mechanism of human glycosyltransferase beta 1,3-N-acetylglucosaminyltransferase 2 (B3GNT2), an important player in immune homeostasis.
Hao, Y., Crequer-Grandhomme, A., Javier, N., Singh, A., Chen, H., Manzanillo, P., Lo, M.C., Huang, X.(2020) J Biological Chem 296: 100042-100042
- PubMed: 33158990 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA120.015306
- Primary Citation Related Structures:
7JHI, 7JHK, 7JHL, 7JHM, 7JHN, 7JHO - PubMed Abstract:
β1,3-N-acetylglucosaminyltransferases (B3GNTs) are Golgi-resident glycosyltransferases involved in the biosynthesis of poly-N-acetyl-lactosamine chains. They catalyze the addition of the N-acetylglucosamine to the N-acetyl-lactosamine repeat as a key step of the chain elongation process. Poly-N-acetyl-lactosamine is involved in the immune system in many ways. Particularly, its long chain has been demonstrated to suppress excessive immune responses. Among the characterized B3GNTs, B3GNT2 is the major poly-N-acetyl-lactosamine synthase, and deletion of its coding gene dramatically reduced the cell surface poly-N-acetyl-lactosamine and led to hypersensitive and hyperresponsive immunocytes. Despite the extensive functional studies, no structural information is available to understand the molecular mechanism of B3GNT2, as well as other B3GNTs. Here we present the structural and kinetic studies of the human B3GNT2. Five crystal structures of B3GNT2 have been determined in the unliganded, donor substrate-bound, acceptor substrate-bound, and product(s)-bound states at resolutions ranging from 1.85 to 2.35 Å. Kinetic study shows that the transglycosylation reaction follows a sequential mechanism. Critical residues involved in recognition of both donor and acceptor substrates as well as catalysis are identified. Mutations of these invariant residues impair B3GNT2 activity in cell assays. Structural comparison with other glycosyltransferases such as mouse Fringe reveals a novel N-terminal helical domain of B3GNTs that may stabilize the catalytic domain and distinguish among different acceptor substrates.
- Department of Molecular Engineering, Amgen Research, Cambridge, Massachusetts, USA; Amgen Postdoctoral Fellow Program, Amgen Research, Cambridge, Massachusetts, USA. Electronic address: yhao@amgen.com.
Organizational Affiliation: