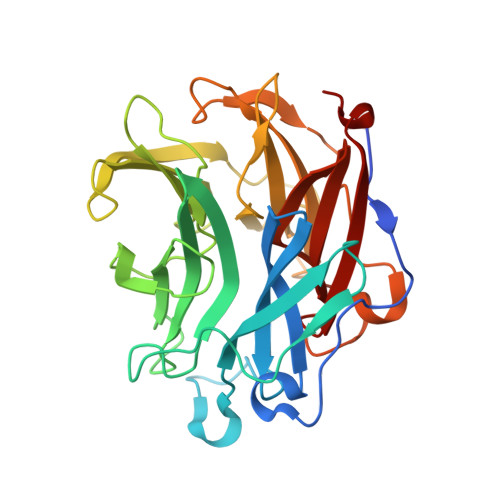
Structural investigation of a thermostable 1,2-beta-mannobiose phosphorylase from Thermoanaerobacter sp. X-514.
Dai, L., Chang, Z., Yang, J., Liu, W., Yang, Y., Chen, C.C., Zhang, L., Huang, J.W., Sun, Y., Guo, R.T.(2021) Biochem Biophys Res Commun 579: 54-61
- PubMed: 34587555 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2021.09.046
- Primary Citation Related Structures:
7FIP, 7FIQ, 7FIR, 7FIS - PubMed Abstract:
1,2-β-Mannobiose phosphorylases (1,2-β-MBPs) from glycoside hydrolase 130 (GH130) family are important bio-catalysts in glycochemistry applications owing to their ability in synthesizing oligomannans. Here, we report the crystal structure of a thermostable 1,2-β-MBP from Thermoanaerobacter sp. X-514 termed Teth514_1789 to reveal the molecular basis of its higher thermostability and mechanism of action. We also solved the enzyme complexes of mannose, mannose-1-phosphate (M1P) and 1,4-β-mannobiose to manifest the enzyme-substrate interaction networks of three main subsites. Notably, a Zn ion that should be derived from crystallization buffer was found in the active site and coordinates the phosphate moiety of M1P. Nonetheless, this Zn-coordination should reflect an inhibitory status as supplementing Zn severely impairs the enzyme activity. These results indicate that the effects of metal ions should be taken into consideration when applying Teth514_1789 and other related enzymes. Based on the structure, a reliable model of Teth514_1788 that shares 61.7% sequence identity to Teth514_1789 but displays a different substrate preference was built. Analyzing the structural features of these two closely related enzymes, we hypothesized that the length of a loop fragment that covers the entrance of the catalytic center might regulate the substrate selectivity. In conclusion, these information provide in-depth understanding of GH130 1,2-β-MBPs and should serve as an important guidance for enzyme engineering for further applications.
- National Engineering Laboratory for Industrial Enzymes, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin, 300308, PR China; University of Chinese Academy of Sciences, Beijing, 100049, PR China; State Key Laboratory of Biocatalysis and Enzyme Engineering, Hubei Collaborative Innovation Center for Green Transformation of Bio-Resources, Hubei Key Laboratory of Industrial Biotechnology, School of Life Sciences, Hubei University, Wuhan, 430062, PR China.
Organizational Affiliation: