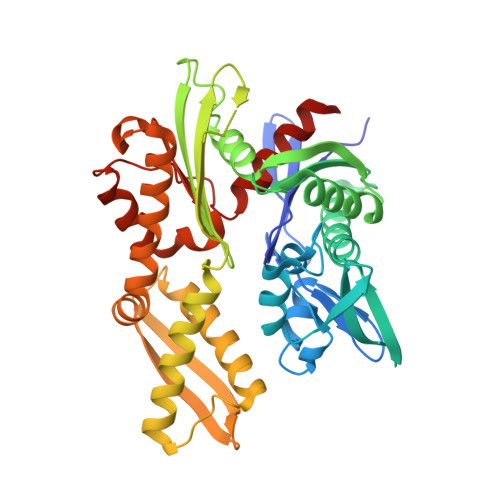
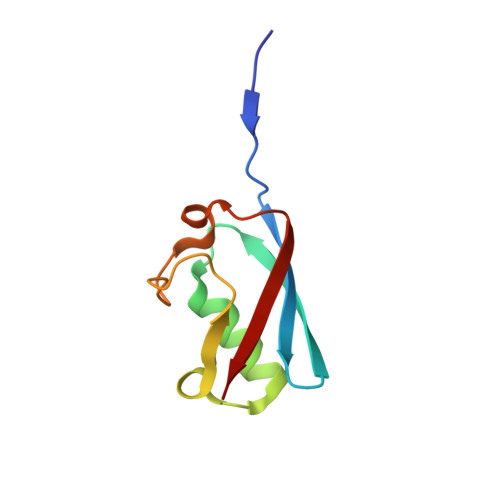
The complex of Fas-associated factor 1 with Hsp70 stabilizes the adherens junction integrity by suppressing RhoA activation
Song, S., Park, J.K., Shin, S.C., Lee, J.J., Hong, S.K., Song, I.K., Kim, B., Song, E.J., Lee, K.J., Kim, E.E.(2022) J Mol Cell Biol 14