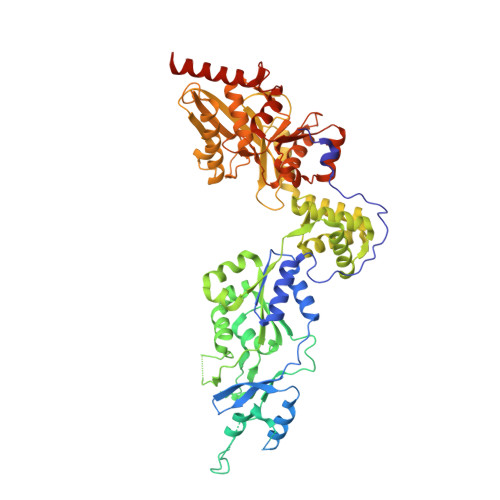
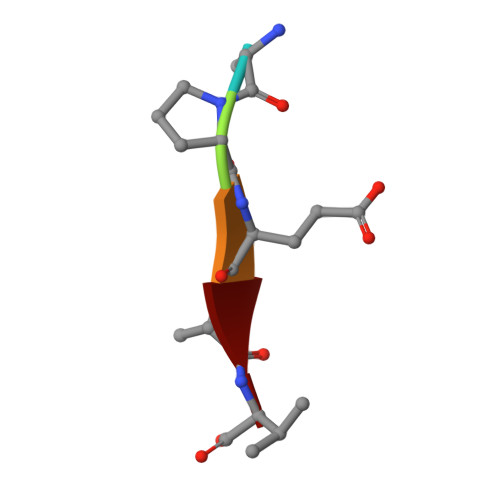
Processive cleavage of substrate at individual proteolytic active sites of the Lon protease complex.
Li, S., Hsieh, K.Y., Kuo, C.I., Su, S.C., Huang, K.F., Zhang, K., Chang, C.I.(2021) Sci Adv 7: eabj9537-eabj9537
- PubMed: 34757797 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abj9537
- Primary Citation Related Structures:
7EUX, 7EUY, 7EV4, 7EV6, 7FID, 7FIE, 7FIZ - PubMed Abstract:
The Lon protease is the prototype of a family of proteolytic machines with adenosine triphosphatase modules built into a substrate degradation chamber. Lon is known to degrade protein substrates in a processive fashion, cutting a protein chain processively into small peptides before commencing cleavages of another protein chain. Here, we present structural and biochemical evidence demonstrating that processive substrate degradation occurs at each of the six proteolytic active sites of Lon, which forms a deep groove that partially encloses the substrate polypeptide chain by accommodating only the unprimed residues and permits processive cleavage in the C-to-N direction. We identify a universally conserved acidic residue at the exit side of the binding groove indispensable for the proteolytic activity. This noncatalytic residue likely promotes processive proteolysis by carboxyl-carboxylate interactions with cleaved intermediates. Together, these results uncover a previously unrecognized mechanism for processive substrate degradation by the Lon protease.
- MOE Key Laboratory for Membraneless Organelles and Cellular Dynamics, Hefei National Laboratory for Physical Sciences at the Microscale and Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei 230027, China.
Organizational Affiliation: