Discovery of pentapeptide-inhibitor hits targeting FKBP51 by combining computational modeling and X-ray crystallography.
Han, J.T., Zhu, Y., Pan, D.B., Xue, H.X., Wang, S., Peng, Y., Liu, H., He, Y.X., Yao, X.(2021) Comput Struct Biotechnol J 19: 4079-4091
- PubMed: 34401048 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.csbj.2021.07.015
- Primary Citation Related Structures:
7ETT, 7ETU, 7ETV - PubMed Abstract:
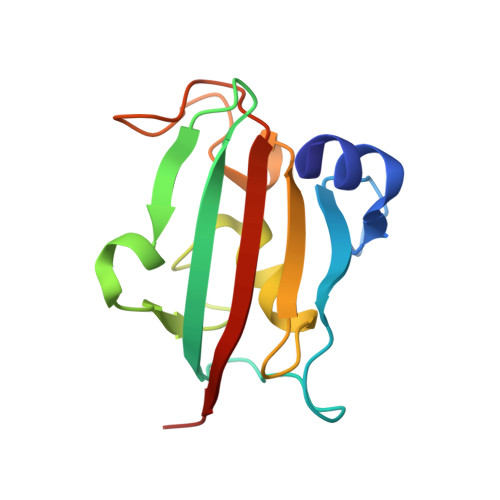
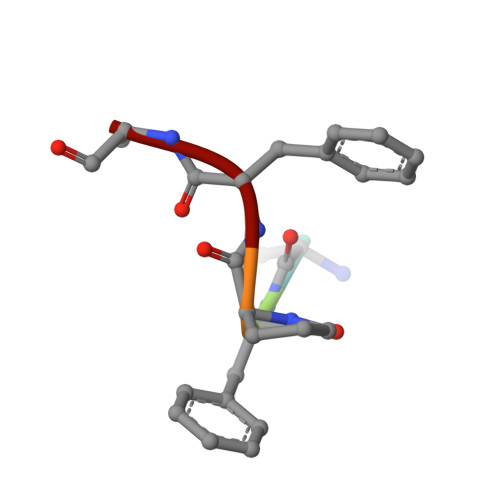
FKBP51 is well-known as a cochaperone of Hsp90 machinery and implicated in many human diseases including stress-related diseases, tau-mediated neurodegeneration and cancers, which makes FKBP51 an attractive drug target for the therapy of FKBP51-associated diseases. However, it has been reported that only nature product rapamycin, cyclosporine A, FK506 and its derivatives exhibit good binding affinities when bound to FKBP51 by now. Given the advantages of peptide-inhibitors, we designed and obtained 20 peptide-inhibitor hits through structure-based drug design. We further characterized the interaction modes of the peptide-inhibitor hits on the FK1 domain of FKBP51 by biochemical and structural biology methods. Structural analysis revealed that peptide-inhibitor hits form U-shaped conformations and occupy the FK506 binding pocket and share similar interaction modes with FK506. Using molecular dynamics simulations, we delved into the interaction dynamics and found that hits are anchored to the FK506 binding pocket in a quite stable conformation. Meanwhile, it was shown that interactions between FK1 and peptide-inhibitor hits are mainly attributed to the hydrogen bond networks comprising I87 and Y113 and FPF cores of peptide-inhibitors involved extensive hydrophobic interactions. We presumed that the peptide design strategy based on the small molecule structure probably shed new lights on the peptide-inhibitor discovery of other targets. The findings presented here could also serve as a structural basis and starting point facilitating the optimization and generation of FKBP51 peptide-inhibitors with better bio-activities.
- State Key Laboratory of Applied Organic Chemistry and Department of Chemistry, Lanzhou University, Lanzhou 730000, China.
Organizational Affiliation: