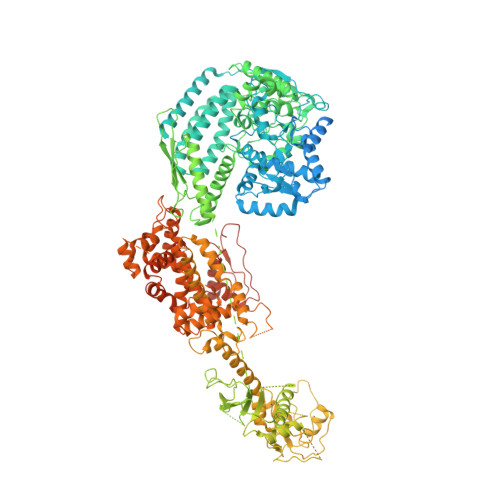
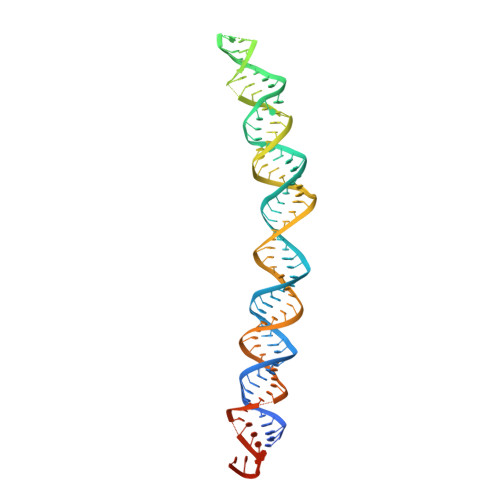
Structural basis of microRNA processing by Dicer-like 1.
Wei, X., Ke, H., Wen, A., Gao, B., Shi, J., Feng, Y.(2021) Nat Plants 7: 1389-1396
- PubMed: 34593993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41477-021-01000-1
- Primary Citation Related Structures:
7ELD, 7ELE - PubMed Abstract:
MicroRNAs (miRNAs) are short non-coding RNAs that inhibit the expression of target genes by directly binding to their mRNAs. In animals, pri-miRNAs are cleaved by Drosha to generate pre-miRNAs, which are subsequently cleaved by Dicer to generate mature miRNAs. Instead of being cleaved by two different enzymes, both cleavages in plants are performed by Dicer-like 1 (DCL1). With a similar domain architecture as human Dicer, it is mysterious how DCL1 recognizes pri-miRNAs and performs two cleavages sequentially. Here, we report the single-particle cryo-electron microscopy structures of Arabidopsis DCL1 complexed with a pri-miRNA and a pre-miRNA, respectively, in cleavage-competent states. These structures uncover the plasticity of the PAZ domain, which is critical for the recognition of both pri-miRNA and pre-miRNA. These structures suggest that the helicase module serves as an engine that transfers the substrate between two sequential cleavage events. This study lays a foundation for dissecting the regulation mechanism of miRNA biogenesis in plants and provides insights into the dicing state of human Dicer.
- Department of Biophysics and Department of Pathology of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, Hangzhou, China.
Organizational Affiliation: