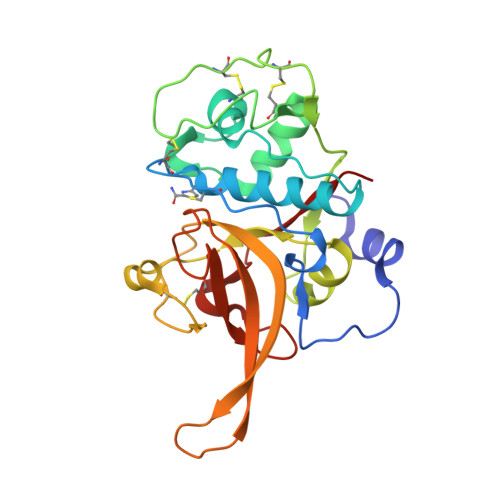
New insights of falcipain 2 structure from Plasmodium falciparum 3D7 strain.
Chakraborty, S., Alam, B., Biswas, S.(2022) Biochem Biophys Res Commun 590: 145-151
- PubMed: 34974303
- DOI: https://doi.org/10.1016/j.bbrc.2021.12.080
- Primary Citation Related Structures:
6JW9, 7EI0 - PubMed Abstract:
Malaria identifies as a tropical hallmark, conforming to the burgeoning notion of escalating drug resistance among virulent strains, with the burdensome Plasmodium falciparum under its wing. The cysteine protease Falcipain-2 (FP2) is released in the parasite's food vacuole in the trophozoite stage and contributes to disease progression through its hemoglobinase activity. In the present study, we have determined the crystal structure of FP2 from a drug resistant P. falciparum 3D7 strain. FP2-3D7 sequence has detected four amino acid variants, R12K, E14 N, P100T and G102D, in the mature domain of the protease, when compared with other reported structures. FP2-3D7 protease has been crystallized in the presence of two inhibitors E-64 and Iodoacetamide, which diffracted up to 3.5 Å and 3.4 Å respectively. Structural analyses of the mature domain helped unveil two solvent-exposed pockets with bound ligands where one is structurally homologous to the allosteric binding site of human Cathepsin-K and thus, could be exploited for designing allosteric modifiers of FP2. The structure has also revealed (poly)ethylene glycol molecules along the catalytic cleft, providing interesting insights for exploring FP2 as a chemotherapeutic target and for PEGylation in drug delivery. The side-chains of P2 and P3 residues of E-64 also adopt different rotamer conformations, compared with previously reported structure, emphasizing strain-specific multiple binding-modes of active-site targeted inhibitors. Docking studies, along with normal mode analyses, highlight the mode of hemoglobin binding and the active/inactive switch in hemoglobinase activity, conjecturing the formation of a stable dimeric state with a symmetry related copy in crystal packing.
- Crystallography and Molecular Biology Division, Saha Institute of Nuclear Physics, HBNI, Kolkata, 700064, India.
Organizational Affiliation: