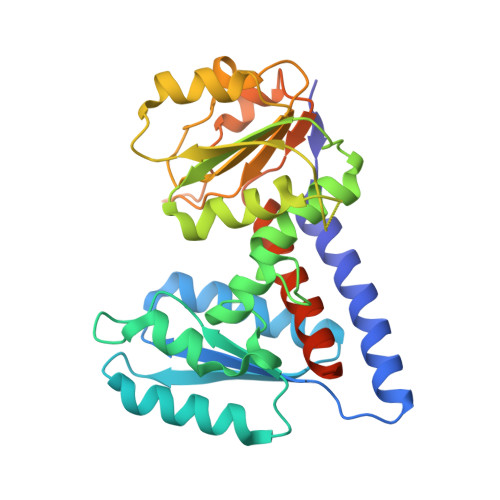
Xanthine Derivatives Reveal an Allosteric Binding Site in Methylenetetrahydrofolate Dehydrogenase 2 (MTHFD2).
Lee, L.C., Peng, Y.H., Chang, H.H., Hsu, T., Lu, C.T., Huang, C.H., Hsueh, C.C., Kung, F.C., Kuo, C.C., Jiaang, W.T., Wu, S.Y.(2021) J Med Chem 64: 11288-11301
- PubMed: 34337952 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.1c00663
- Primary Citation Related Structures:
7EHJ, 7EHM, 7EHN, 7EHV - PubMed Abstract:
Methylenetetrahydrofolate dehydrogenase 2 (MTHFD2) plays an important role in one-carbon metabolism. The MTHFD2 gene is upregulated in various cancers but very low or undetectable in normal proliferating cells, and therefore a potential target for cancer treatment. In this study, we present the structure of MTHFD2 in complex with xanthine derivative 15 , which allosterically binds to MTHFD2 and coexists with the substrate analogue. A kinetic study demonstrated the uncompetitive inhibition of MTHFD2 by 15 . Allosteric inhibitors often provide good selectivity and, indeed, xanthine derivatives are highly selective for MTHFD2. Moreover, several conformational changes were observed upon the binding of 15 , which impeded the binding of the cofactor and phosphate to MTHFD2. To the best of our knowledge, this is the first study to identify allosteric inhibitors targeting the MTHFD family and our results would provide insights on the inhibition mechanism of MTHFD proteins and the development of novel inhibitors.
- Institute of Biotechnology and Pharmaceutical Research, National Health Research Institutes, 35, Keyan Road, Zhunan Town, Miaoli County 350 Taiwan, ROC.
Organizational Affiliation: