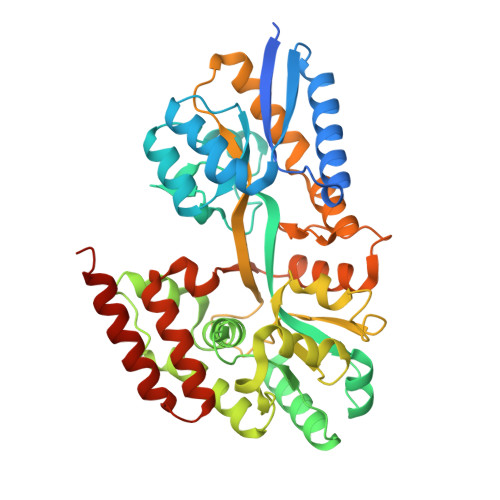
Structural characterization of two solute-binding proteins for N,N' -diacetylchitobiose/ N,N',N'' -triacetylchitotoriose of the gram-positive bacterium, Paenibacillus sp. str. FPU-7.
Itoh, T., Yaguchi, M., Nakaichi, A., Yoda, M., Hibi, T., Kimoto, H.(2021) J Struct Biol X 5: 100049-100049
- PubMed: 34195603 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.yjsbx.2021.100049
- Primary Citation Related Structures:
7EHO, 7EHP, 7EHQ, 7EHU - PubMed Abstract:
The chitinolytic bacterium Paenibacillus sp. str. FPU-7 efficiently degrades chitin into oligosaccharides such as N -acetyl-D-glucosamine (GlcNAc) and disaccharides (GlcNAc) 2 through multiple secretory chitinases. Transport of these oligosaccharides by P . str. FPU-7 has not yet been clarified. In this study, we identified nagB1 , predicted to encode a sugar solute-binding protein (SBP), which is a component of the ABC transport system. However, the genes next to nagB1 were predicted to encode two-component regulatory system proteins rather than transmembrane domains (TMDs). We also identified nagB2 , which is highly homologous to nagB1 . Adjacent to nagB2 , two genes were predicted to encode TMDs. Binding experiments of the recombinant NagB1 and NagB2 to several oligosaccharides using differential scanning fluorimetry and surface plasmon resonance confirmed that both proteins are SBPs of (GlcNAc) 2 and (GlcNAc) 3 . We determined their crystal structures complexed with and without chitin oligosaccharides at a resolution of 1.2 to 2.0 Å. The structures shared typical SBP structural folds and were classified as subcluster D-I. Large domain motions were observed in the structures, suggesting that they were induced by ligand binding via the "Venus flytrap" mechanism. These structures also revealed chitin oligosaccharide recognition mechanisms. In conclusion, our study provides insight into the recognition and transport of chitin oligosaccharides in bacteria.
- Department of Bioscience and Biotechnology, Fukui Prefectural University, 4-1-1 Matsuokakenjyoujima, Eiheiji-cho, Yoshida-gun, Fukui 910-1195, Japan.
Organizational Affiliation: