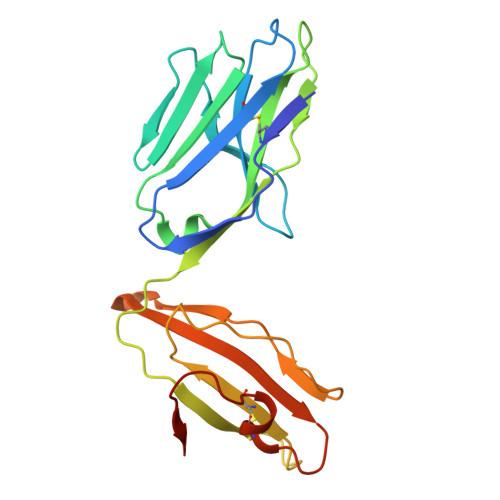
Identification of conserved SARS-CoV-2 spike epitopes that expand public cTfh clonotypes in mild COVID-19 patients.
Lu, X., Hosono, Y., Nagae, M., Ishizuka, S., Ishikawa, E., Motooka, D., Ozaki, Y., Sax, N., Maeda, Y., Kato, Y., Morita, T., Shinnakasu, R., Inoue, T., Onodera, T., Matsumura, T., Shinkai, M., Sato, T., Nakamura, S., Mori, S., Kanda, T., Nakayama, E.E., Shioda, T., Kurosaki, T., Takeda, K., Kumanogoh, A., Arase, H., Nakagami, H., Yamashita, K., Takahashi, Y., Yamasaki, S.(2021) J Exp Medicine 218
- PubMed: 34647971 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1084/jem.20211327
- Primary Citation Related Structures:
7EA6 - PubMed Abstract:
Adaptive immunity is a fundamental component in controlling COVID-19. In this process, follicular helper T (Tfh) cells are a subset of CD4+ T cells that mediate the production of protective antibodies; however, the SARS-CoV-2 epitopes activating Tfh cells are not well characterized. Here, we identified and crystallized TCRs of public circulating Tfh (cTfh) clonotypes that are expanded in patients who have recovered from mild symptoms. These public clonotypes recognized the SARS-CoV-2 spike (S) epitopes conserved across emerging variants. The epitope of the most prevalent cTfh clonotype, S864-882, was presented by multiple HLAs and activated T cells in most healthy donors, suggesting that this S region is a universal T cell epitope useful for booster antigen. SARS-CoV-2-specific public cTfh clonotypes also cross-reacted with specific commensal bacteria. In this study, we identified conserved SARS-CoV-2 S epitopes that activate public cTfh clonotypes associated with mild symptoms.
- Laboratory of Molecular Immunology, Immunology Frontier Research Center, Osaka University, Suita, Japan.
Organizational Affiliation: