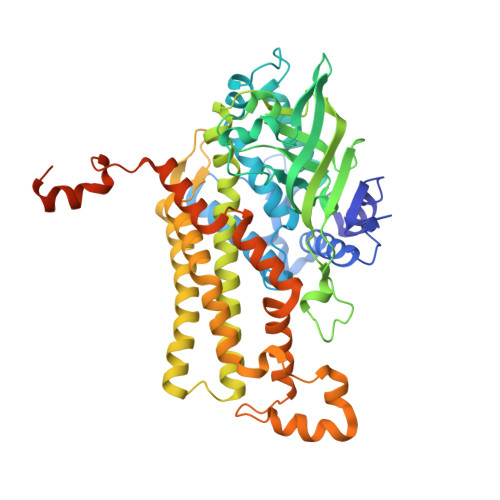
Structural insights into a flavin-dependent dehalogenase HadA explain catalysis and substrate inhibition via quadruple pi-stacking.
Pimviriyakul, P., Jaruwat, A., Chitnumsub, P., Chaiyen, P.(2021) J Biological Chem 297: 100952-100952
- PubMed: 34252455 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jbc.2021.100952
- Primary Citation Related Structures:
7E8P, 7E8Q - PubMed Abstract:
HadA is a flavin-dependent monooxygenase catalyzing hydroxylation plus dehalogenation/denitration, which is useful for biodetoxification and biodetection. In this study, the X-ray structure of wild-type HadA (HadA WT ) co-complexed with reduced FAD (FADH - ) and 4-nitrophenol (4NP) (HadA WT -FADH - -4NP) was solved at 2.3-Å resolution, providing the first full package (with flavin and substrate bound) structure of a monooxygenase of this type. Residues Arg101, Gln158, Arg161, Thr193, Asp254, Arg233, and Arg439 constitute a flavin-binding pocket, whereas the 4NP-binding pocket contains the aromatic side chain of Phe206, which provides π-π stacking and also is a part of the hydrophobic pocket formed by Phe155, Phe286, Thr449, and Leu457. Based on site-directed mutagenesis and stopped-flow experiments, Thr193, Asp254, and His290 are important for C4a-hydroperoxyflavin formation with His290, also serving as a catalytic base for hydroxylation. We also identified a novel structural motif of quadruple π-stacking (π-π-π-π) provided by two 4NP and two Phe441 from two subunits. This motif promotes 4NP binding in a nonproductive dead-end complex, which prevents C4a-hydroperoxy-FAD formation when HadA is premixed with aromatic substrates. We also solved the structure of the HadA Phe441Val -FADH - -4NP complex at 2.3-Å resolution. Although 4NP can still bind to this variant, the quadruple π-stacking motif was disrupted. All HadA Phe441 variants lack substrate inhibition behavior, confirming that quadruple π-stacking is a main cause of dead-end complex formation. Moreover, the activities of these HadA Phe441 variants were improved by ⁓20%, suggesting that insights gained from the flavin-dependent monooxygenases illustrated here should be useful for future improvement of HadA's biocatalytic applications.
- Department of Biochemistry, Faculty of Science, Kasetsart University, Bangkok, Thailand.
Organizational Affiliation: