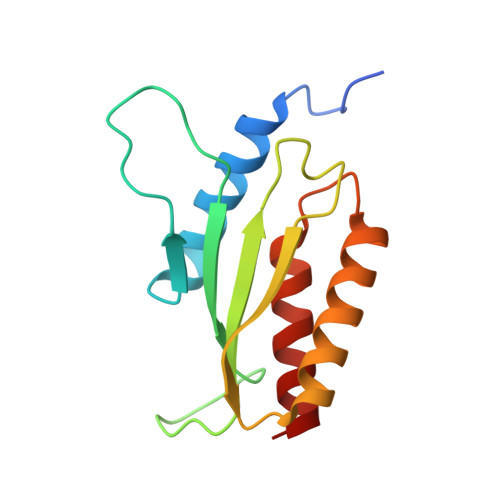
Crystal structures reveal a novel dimer of the RWD domain of human general control nonderepressible 2.
Hei, Z., Wu, S., Zheng, L., Zhou, J., Liu, Z., Wang, J., Fang, P.(2021) Biochem Biophys Res Commun 549: 164-170
- PubMed: 33676185 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2021.02.111
- Primary Citation Related Structures:
7E2K, 7E2M - PubMed Abstract:
General control nonderepressible 2 (GCN2) is a serine/threonine protein kinase, detecting a variety of stresses including amino acid starvation, reactive oxygen species, etc. in eukaryotic cells. Activation of GCN2 requires the interaction of the N-terminal RWD domain with the upstream GCN1 protein and the dimerization by the kinase domain. In this study, we determined two crystal structures of the RWD domain of human GCN2 in two different crystal packing modes. These two different crystal structures reveal a same dimer of the RWD domain, which has not been reported in previous studies. We further confirmed this novel dimer interaction in solution using gel filtration experiments, and in human embryonic kidney (HEK) 293 cells using bimolecular fluorescence complementation (BiFC) and co-immunoprecipitation (co-IP) assays. Together, this study discovers a potential protein-protein interface on the RWD domain of human GCN2, and suggests a possible regulation between the interaction of GCN1 and the formation of GCN2 dimer.
- State Key Laboratory of Bioorganic and Natural Products Chemistry, Center for Excellence in Molecular Synthesis, Shanghai Institute of Organic Chemistry, University of Chinese Academy of Sciences, Chinese Academy of Sciences, 345 Lingling Road, Shanghai, 200032, China.
Organizational Affiliation: