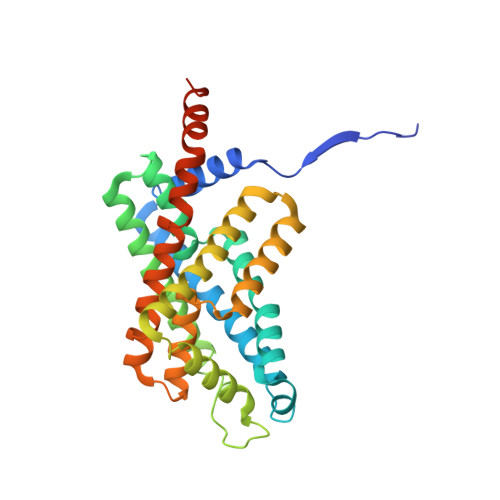
Structural characterization of the Plasmodium falciparum lactate transporter PfFNT alone and in complex with antimalarial compound MMV007839 reveals its inhibition mechanism.
Peng, X., Wang, N., Zhu, A., Xu, H., Li, J., Zhou, Y., Wang, C., Xiao, Q., Guo, L., Liu, F., Jia, Z.J., Duan, H., Hu, J., Yuan, W., Geng, J., Yan, C., Jiang, X., Deng, D.(2021) PLoS Biol 19: e3001386-e3001386
- PubMed: 34499638 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.3001386
- Primary Citation Related Structures:
7E26, 7E27 - PubMed Abstract:
Plasmodium falciparum, the deadliest causal agent of malaria, caused more than half of the 229 million malaria cases worldwide in 2019. The emergence and spreading of frontline drug-resistant Plasmodium strains are challenging to overcome in the battle against malaria and raise urgent demands for novel antimalarial agents. The P. falciparum formate-nitrite transporter (PfFNT) is a potential drug target due to its housekeeping role in lactate efflux during the intraerythrocytic stage. Targeting PfFNT, MMV007839 was identified as a lead compound that kills parasites at submicromolar concentrations. Here, we present 2 cryogenic-electron microscopy (cryo-EM) structures of PfFNT, one with the protein in its apo form and one with it in complex with MMV007839, both at 2.3 Å resolution. Benefiting from the high-resolution structures, our study provides the molecular basis for both the lactate transport of PfFNT and the inhibition mechanism of MMV007839, which facilitates further antimalarial drug design.
- Department of Obstetrics, Key Laboratory of Birth Defects and Related Disease of Women and Children of MOE, State Key Laboratory of Biotherapy, West China Second Hospital, Sichuan University, Chengdu, China.
Organizational Affiliation: