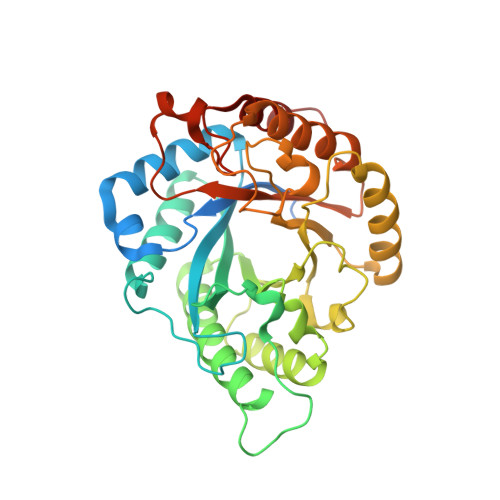
Functional and structural investigation of a novel beta-mannanase BaMan113A from Bacillus sp. N16-5.
Liu, W., Ma, C., Liu, W., Zheng, Y., Chen, C.C., Liang, A., Luo, X., Li, Z., Ma, W., Song, Y., Guo, R.T., Zhang, T.(2021) Int J Biol Macromol 182: 899-909
- PubMed: 33865894 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2021.04.075
- Primary Citation Related Structures:
7DV7, 7DVJ, 7DVZ, 7DW8, 7DWA - PubMed Abstract:
Mannan is an important renewable resource whose backbone can be hydrolyzed by β-mannanases to generate manno-oligosaccharides of various sizes. Only a few glycoside hydrolase (GH) 113 family β-mannanases have been functionally and structurally characterize. Here, we report the function and structure of a novel GH113 β-mannanase from Bacillus sp. N16-5 (BaMan113A). BaMan113A exhibits a substrate preference toward manno-oligosaccharides and releases mannose and mannobiose as main hydrolytic products. The crystal structure of BaMan113A suggest that the enzyme shows a semi-enclosed substrate-binding cleft and the amino acids surrounding the +2 subsite form a steric barrier to terminate the substrate-binding tunnel. Based on these structural features, we conducted mutagenesis to engineer BaMan113A to remove the steric hindrance of the substrate-binding tunnel. We found that F101E and N236Y variants exhibit increased specific activity toward mannans comparing to the wild-type enzyme. Meanwhile, the product profiles of these two variants toward polysaccharides changed from mannose to a series of manno-oligosaccharides. The crystal structure of variant N236Y was also determined to illustrate the molecular basis underlying the mutation. In conclusion, we report the functional and structural features of a novel GH113 β-mannanase, and successfully improved its endo-acting activity by using structure-based engineering.
- Key Laboratory of Industrial Fermentation Microbiology of Ministry of Education & Tianjin Key Laboratory of Industrial Microbiology, College of Biotechnology, Tianjin University of Science and Technology, Tianjin 300457, China; Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin 300308, China.
Organizational Affiliation: