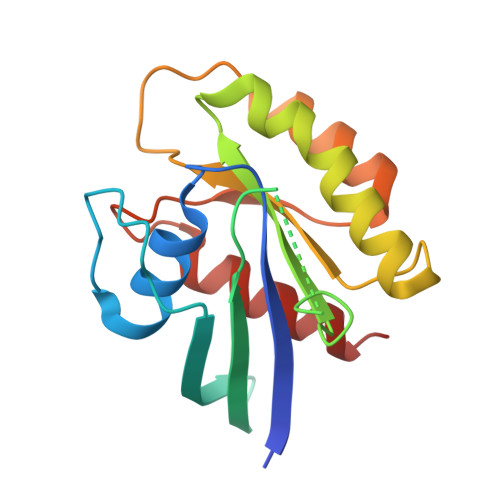
Oncogenic mutations Q61L and Q61H confer active form-like structural features to the inactive state (state 1) conformation of H-Ras protein.
Matsumoto, S., Taniguchi-Tamura, H., Araki, M., Kawamura, T., Miyamoto, R., Tsuda, C., Shima, F., Kumasaka, T., Okuno, Y., Kataoka, T.(2021) Biochem Biophys Res Commun 565: 85-90
- PubMed: 34102474 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2021.05.084
- Primary Citation Related Structures:
7DPH, 7DPJ - PubMed Abstract:
GTP-bound forms of Ras proteins (Ras•GTP) assume two interconverting conformations, "inactive" state 1 and "active" state 2. Our previous study on the crystal structure of the state 1 conformation of H-Ras in complex with guanosine 5'-(β, γ-imido)triphosphate (GppNHp) indicated that state 1 is stabilized by intramolecular hydrogen-bonding interactions formed by Gln61. Since Ras are constitutively activated by substitution mutations of Gln61, here we determine crystal structures of the state 1 conformation of H-Ras•GppNHp carrying representative mutations Q61L and Q61H to observe the effect of the mutations. The results show that these mutations alter the mode of hydrogen-bonding interactions of the residue 61 with Switch II residues and induce conformational destabilization of the neighboring regions. In particular, Q61L mutation results in acquirement of state 2-like structural features. Moreover, the mutations are likely to impair an intramolecular structural communication between Switch I and Switch II. Molecular dynamics simulations starting from these structures support the above observations. These findings may give a new insight into the molecular mechanism underlying the aberrant activation of the Gln61 mutants.
- Department of Biomedical Data Intelligence, Graduate School of Medicine, Kyoto University, 53 Shogoin-Kawaharacho, Sakyo-ku, Kyoto, 606-8507, Japan.
Organizational Affiliation: