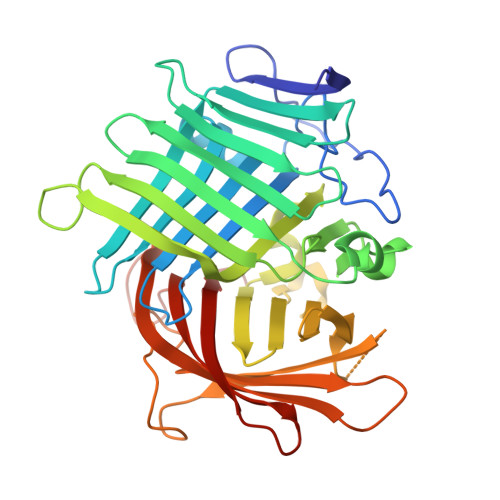
Crystal Structures of Fsa2 and Phm7 Catalyzing [4 + 2] Cycloaddition Reactions with Reverse Stereoselectivities in Equisetin and Phomasetin Biosynthesis.
Chi, C., Wang, Z., Liu, T., Zhang, Z., Zhou, H., Li, A., Jin, H., Jia, H., Yin, F., Yang, D., Ma, M.(2021) ACS Omega 6: 12913-12922
- PubMed: 34056443 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsomega.1c01593
- Primary Citation Related Structures:
7DMN, 7DMO - PubMed Abstract:
Fsa2 and Phm7 are a unique pair of pericyclases catalyzing [4 + 2] cycloaddition reactions with reverse stereoselectivities in the biosynthesis of equisetin and phomasetin, both of which are potent HIV-1 integrase inhibitors. We here solve the crystal structures of Fsa2 and Phm7, both of which possess unusual "two-β barrel" folds. Different residues are evident between the active sites of Fsa2 and Phm7, and modeling experiments provide key structural information determining the reverse stereoselectivities. These results provide a better understanding of how natural pericyclases control the catalytic stereoselectivities and benefit the protein engineering in future.
- State Key Laboratory of Natural and Biomimetic Drugs, School of Pharmaceutical Sciences, Peking University, 38 Xueyuan Road, Haidian District, Beijing 100191, China.
Organizational Affiliation: