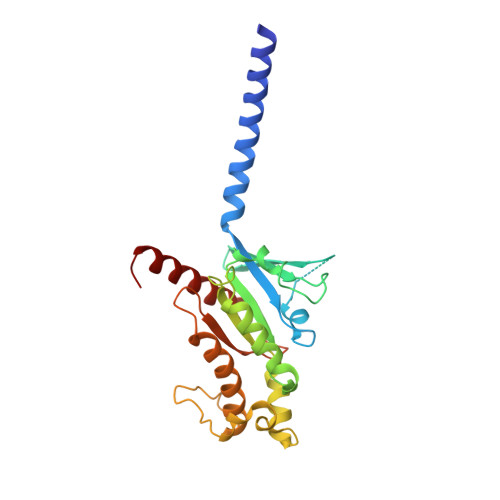
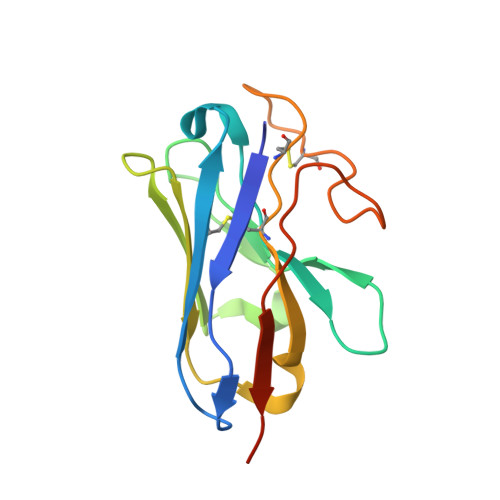
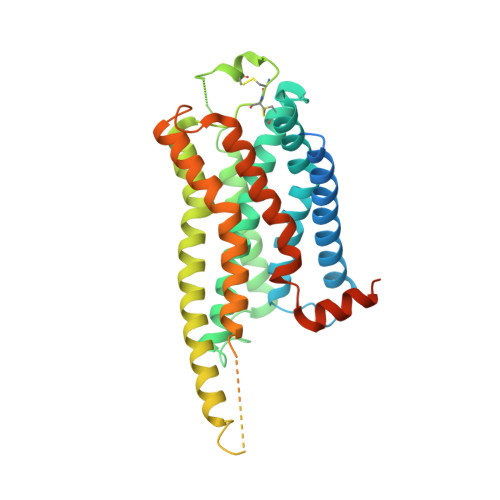
Cryo-EM structure of the beta 3-adrenergic receptor reveals the molecular basis of subtype selectivity.
Nagiri, C., Kobayashi, K., Tomita, A., Kato, M., Kobayashi, K., Yamashita, K., Nishizawa, T., Inoue, A., Shihoya, W., Nureki, O.(2021) Mol Cell 81: 3205-3215.e5
- PubMed: 34314699 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2021.06.024
- Primary Citation Related Structures:
7DH5, 7XJH - PubMed Abstract:
The β 3 -adrenergic receptor (β 3 AR) is predominantly expressed in adipose tissue and urinary bladder and has emerged as an attractive drug target for the treatment of type 2 diabetes, obesity, and overactive bladder (OAB). Here, we report the cryogenic electron microscopy structure of the β 3 AR-G s signaling complex with the selective agonist mirabegron, a first-in-class drug for OAB. Comparison of this structure with the previously reported β 1 AR and β 2 AR structures reveals a receptor activation mechanism upon mirabegron binding to the orthosteric site. Notably, the narrower exosite in β 3 AR creates a perpendicular pocket for mirabegron. Mutational analyses suggest that a combination of both the exosite shape and the amino-acid-residue substitutions defines the drug selectivity of the βAR agonists. Our findings provide a molecular basis for βAR subtype selectivity, allowing the design of more-selective agents with fewer adverse effects.
- Department of Biological Sciences, Graduate School of Science, the University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: