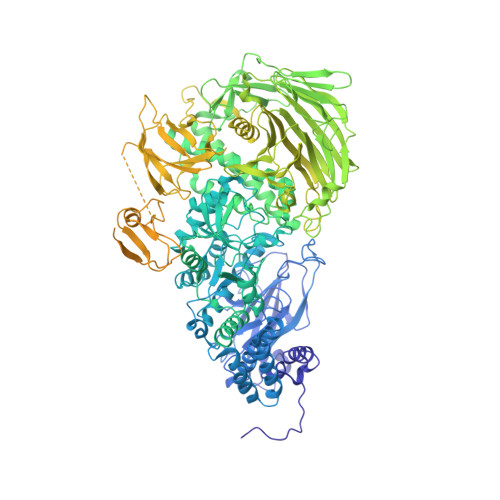
Molecular and structural mechanisms of ZZ domain-mediated cargo selection by Nbr1.
Wang, Y.Y., Zhang, J., Liu, X.M., Li, Y., Sui, J., Dong, M.Q., Ye, K., Du, L.L.(2021) EMBO J 40: e107497-e107497
- PubMed: 34169534 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2020107497
- Primary Citation Related Structures:
7DD9, 7DDE - PubMed Abstract:
In selective autophagy, cargo selectivity is determined by autophagy receptors. However, it remains scarcely understood how autophagy receptors recognize specific protein cargos. In the fission yeast Schizosaccharomyces pombe, a selective autophagy pathway termed Nbr1-mediated vacuolar targeting (NVT) employs Nbr1, an autophagy receptor conserved across eukaryotes including humans, to target cytosolic hydrolases into the vacuole. Here, we identify two new NVT cargos, the mannosidase Ams1 and the aminopeptidase Ape4, that bind competitively to the first ZZ domain of Nbr1 (Nbr1-ZZ1). High-resolution cryo-EM analyses reveal how a single ZZ domain recognizes two distinct protein cargos. Nbr1-ZZ1 not only recognizes the N-termini of cargos via a conserved acidic pocket, similar to other characterized ZZ domains, but also engages additional parts of cargos in a cargo-specific manner. Our findings unveil a single-domain bispecific mechanism of autophagy cargo recognition, elucidate its underlying structural basis, and expand the understanding of ZZ domain-mediated protein-protein interactions.
- College of Life Sciences, Beijing Normal University, Beijing, China.
Organizational Affiliation: