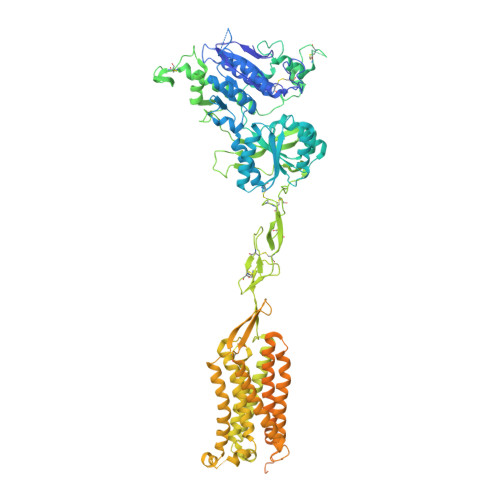
Structural basis for activation and allosteric modulation of full-length calcium-sensing receptor.
Wen, T., Wang, Z., Chen, X., Ren, Y., Lu, X., Xing, Y., Lu, J., Chang, S., Zhang, X., Shen, Y., Yang, X.(2021) Sci Adv 7
- PubMed: 34088669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abg1483
- Primary Citation Related Structures:
7DD5, 7DD6, 7DD7 - PubMed Abstract:
Calcium-sensing receptor (CaSR) is a class C G protein-coupled receptor (GPCR) that plays an important role in calcium homeostasis and parathyroid hormone secretion. Here, we present multiple cryo-electron microscopy structures of full-length CaSR in distinct ligand-bound states. Ligands (Ca 2+ and l-tryptophan) bind to the extracellular domain of CaSR and induce large-scale conformational changes, leading to the closure of two heptahelical transmembrane domains (7TMDs) for activation. The positive modulator (evocalcet) and the negative allosteric modulator (NPS-2143) occupy the similar binding pocket in 7TMD. The binding of NPS-2143 causes a considerable rearrangement of two 7TMDs, forming an inactivated TM6/TM6 interface. Moreover, a total of 305 disease-causing missense mutations of CaSR have been mapped to the structure in the active state, creating hotspot maps of five clinical endocrine disorders. Our results provide a structural framework for understanding the activation, allosteric modulation mechanism, and disease therapy for class C GPCRs.
- State Key Laboratory of Medicinal Chemical Biology and College of Life Sciences, Nankai University, Tianjin 300350, China.
Organizational Affiliation: