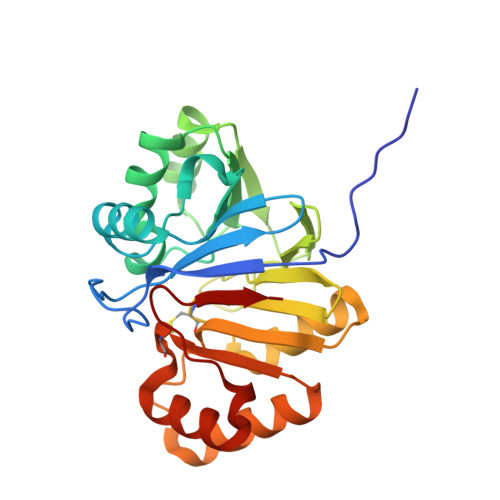
Structural and functional identification of the uncharacterized metallo-beta-lactamase superfamily protein TW9814 as a phosphodiesterase with unique metal coordination.
Heo, Y., Park, S.B., Jeon, Y.E., Yun, J.H., Jeong, B.G., Cha, S.S., Lee, W.(2022) Acta Crystallogr D Struct Biol 78: 532-541
- PubMed: 35362475 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798322002108
- Primary Citation Related Structures:
7DCK - PubMed Abstract:
Metallo-β-lactamase (MBL) superfamily proteins have a common αβ/βα sandwich fold and perform a variety of functions through metal-mediated catalysis. However, because of the enormous scale of this superfamily, only a small percentage of the proteins belonging to the superfamily have been annotated structurally or functionally to date. Therefore, much remains unknown about the MBL superfamily proteins. Here, TW9814, a hypothetical MBL superfamily protein, was structurally and functionally investigated. Guided by the crystal structure of dimeric TW9814, it was demonstrated that TW9814 functions as a phosphodiesterase (PDE) in the presence of divalent metal ions such as manganese(II) or nickel(II). A docking model between TW9814 and the substrate bis(p-nitrophenyl)phosphate (bpNPP) showed the importance of the dimerization of TW9814 for its bpNPP-hydrolyzing activity and for the interaction between the enzyme and the substrate. TW9814 showed outstanding catalytic efficiency (k cat /K m ) under alkaline conditions compared with other PDEs. The activity of TW9814 appears to be regulated through a disulfide bond, which is a feature that is not present in other MBL superfamily members. This study provides a platform for the functional characterization of other hypothetical proteins of the MBL or other superfamilies.
- Department of Biochemistry, Yonsei University, 50 Yonsei-ro, Seoul 03722, Republic of Korea.
Organizational Affiliation: