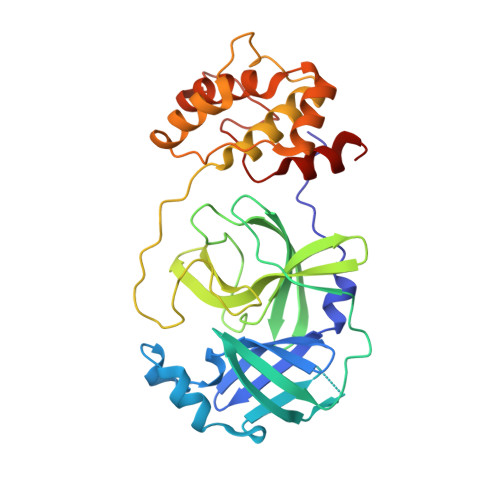
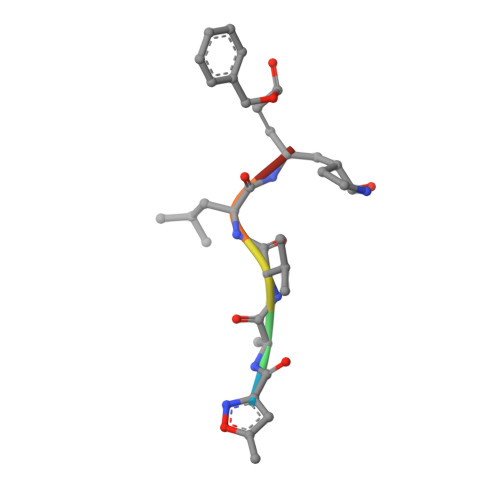
The newly emerged SARS-like coronavirus HCoV-EMC also has an "Achilles' heel": current effective inhibitor targeting a 3C-like protease
Ren, Z., Yan, L., Zhang, N., Guo, Y., Yang, C., Lou, Z., Rao, Z.(2013) Protein Cell 4: 248-250
- PubMed: 23549610 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-013-2841-3
- Primary Citation Related Structures:
7D3C - College of Pharmacy and State Key Laboratory of Medicinal Chemical Biology, Nankai University, Tianjin 300071, China.
Organizational Affiliation: