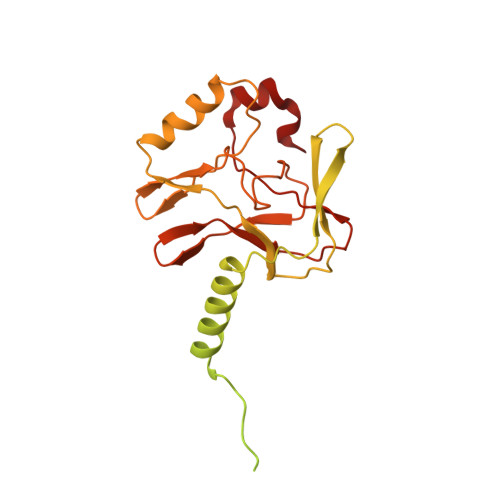
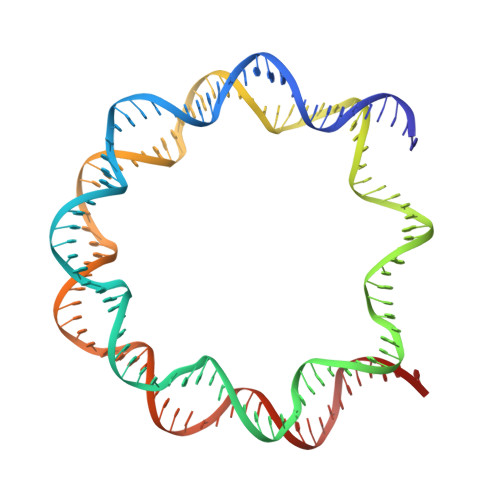
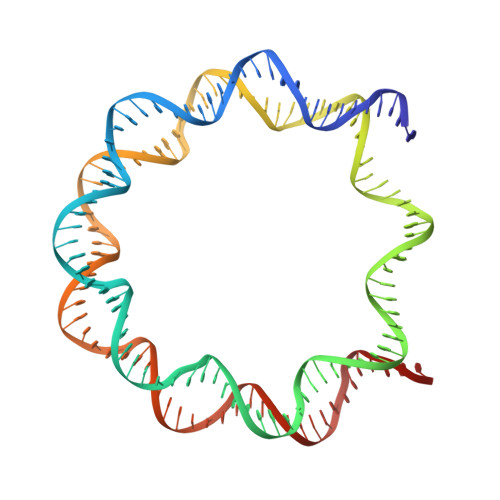
Structural basis of nucleosomal histone H4 lysine 20 methylation by SET8 methyltransferase.
Ho, C.H., Takizawa, Y., Kobayashi, W., Arimura, Y., Kimura, H., Kurumizaka, H.(2021) Life Sci Alliance 4
- PubMed: 33574035 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.26508/lsa.202000919
- Primary Citation Related Structures:
7D1Z, 7D20 - PubMed Abstract:
SET8 is solely responsible for histone H4 lysine-20 (H4K20) monomethylation, which preferentially occurs in nucleosomal H4. However, the underlying mechanism by which SET8 specifically promotes the H4K20 monomethylation in the nucleosome has not been elucidated. Here, we report the cryo-EM structures of the human SET8-nucleosome complexes with histone H3 and the centromeric H3 variant, CENP-A. Surprisingly, we found that the overall cryo-EM structures of the SET8-nucleosome complexes are substantially different from the previous crystal structure models. In the complexes with H3 and CENP-A nucleosomes, SET8 specifically binds the nucleosomal acidic patch via an arginine anchor, composed of the Arg188 and Arg192 residues. Mutational analyses revealed that the interaction between the SET8 arginine anchor and the nucleosomal acidic patch plays an essential role in the H4K20 monomethylation activity. These results provide the groundwork for understanding the mechanism by which SET8 specifically accomplishes the H4K20 monomethylation in the nucleosome.
- Laboratory of Chromatin Structure and Function, Institute for Quantitative Biosciences, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: