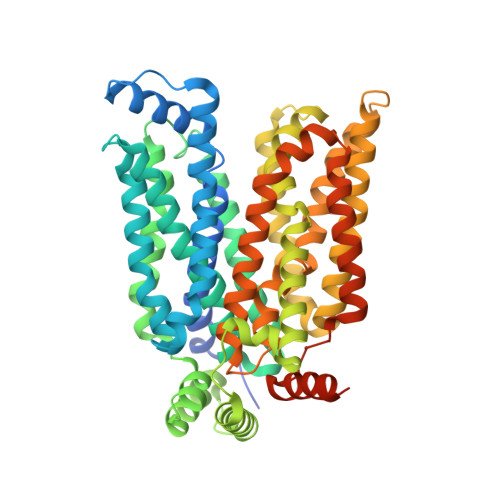
Orthosteric-allosteric dual inhibitors of PfHT1 as selective antimalarial agents.
Huang, J., Yuan, Y., Zhao, N., Pu, D., Tang, Q., Zhang, S., Luo, S., Yang, X., Wang, N., Xiao, Y., Zhang, T., Liu, Z., Sakata-Kato, T., Jiang, X., Kato, N., Yan, N., Yin, H.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33402433 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2017749118
- Primary Citation Related Structures:
7CRZ - PubMed Abstract:
Artemisinin-resistant malaria parasites have emerged and have been spreading, posing a significant public health challenge. Antimalarial drugs with novel mechanisms of action are therefore urgently needed. In this report, we exploit a "selective starvation" strategy by inhibiting Plasmodium falciparum hexose transporter 1 (PfHT1), the sole hexose transporter in P. falciparum , over human glucose transporter 1 (hGLUT1), providing an alternative approach to fight against multidrug-resistant malaria parasites. The crystal structure of hGLUT3, which shares 80% sequence similarity with hGLUT1, was resolved in complex with C3361, a moderate PfHT1-specific inhibitor, at 2.3-Å resolution. Structural comparison between the present hGLUT3-C3361 and our previously reported PfHT1-C3361 confirmed the unique inhibitor binding-induced pocket in PfHT1. We then designed small molecules to simultaneously block the orthosteric and allosteric pockets of PfHT1. Through extensive structure-activity relationship studies, the TH-PF series was identified to selectively inhibit PfHT1 over hGLUT1 and potent against multiple strains of the blood-stage P. falciparum Our findings shed light on the next-generation chemotherapeutics with a paradigm-shifting structure-based design strategy to simultaneously target the orthosteric and allosteric sites of a transporter.
- Key Laboratory of Bioorganic Phosphorous Chemistry and Chemical Biology (Ministry of Education), Department of Chemistry, School of Pharmaceutical Sciences, Tsinghua University,100084 Beijing, China.
Organizational Affiliation: