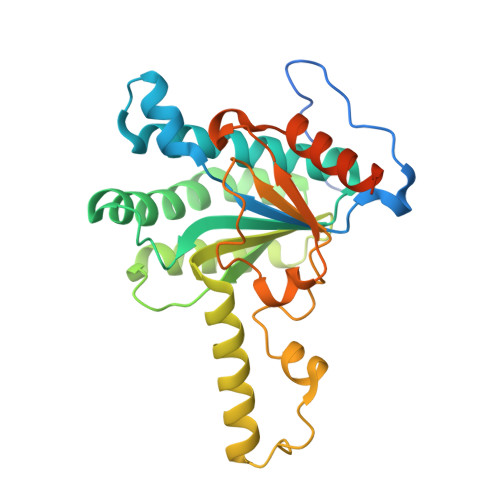
Crystal structure of Thermobifida fusca cis-prenyltransferase reveals the dynamic nature of its RXG motif-mediated inter-subunit interactions critical for its catalytic activity.
Kurokawa, H., Ambo, T., Takahashi, S., Koyama, T.(2020) Biochem Biophys Res Commun 532: 459-465
- PubMed: 32892948 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.08.062
- Primary Citation Related Structures:
7CPM, 7CPN - PubMed Abstract:
cis-Prenyltransferases (cis-PTs) catalyze consecutive condensations of isopentenyl diphosphate to an allylic diphosphate acceptor to produce a linear polyprenyl diphosphate of designated length. Dimer formation is a prerequisite for cis-PTs to catalyze all cis-prenyl condensation reactions. The structure-function relationship of a conserved C-terminal RXG motif in cis-PTs that forms inter-subunit interactions and has a role in catalytic activity has attracted much attention. Here, we solved the crystal structure of a medium-chain cis-PT from Thermobifida fusca that produces dodecaprenyl diphosphate as a polyprenoid glycan carrier for cell wall synthesis. The structure revealed a characteristic dimeric architecture of cis-PTs in which a rigidified RXG motif of one monomer formed inter-subunit hydrogen bonds with the catalytic site of the other monomer, while the RXG motif of the latter remained flexible. Careful analyses suggested the existence of a possible long-range negative cooperativity between the two catalytic sites on the two monomeric subunits that allowed the binding of one subunit to stabilize the formation of the enzyme-substrate ternary complex and facilitated the release of Mg-PPi and subsequent intra-molecular translocation at the counter subunit so that the condensation reaction could occur in consecutive cycles. The current structure reveals the dynamic nature of the RXG motif and provides a rationale for pursuing further investigations to elucidate the inter-subunit cooperativity of cis-PTs.
- Institute of Multidisciplinary Research for Advanced Materials, Tohoku University, Katahira 2-1-1, Aoba-ku, Sendai, 980-8577, Japan. Electronic address: hirofumi.kurokawa.e2@tohoku.ac.jp.
Organizational Affiliation: