Structural characterization of a neutralizing mAb H16.001, a potent candidate for a common potency assay for various HPV16 VLPs.
Huang, W., He, M., Ning, T., Nie, J., Zhang, F., Zheng, Q., Zhang, R., Xu, Y., Gu, Y., Li, S., Wang, Y.(2020) NPJ Vaccines 5: 89-89
- PubMed: 33042588 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41541-020-00236-w
- Primary Citation Related Structures:
7CN2 - PubMed Abstract:
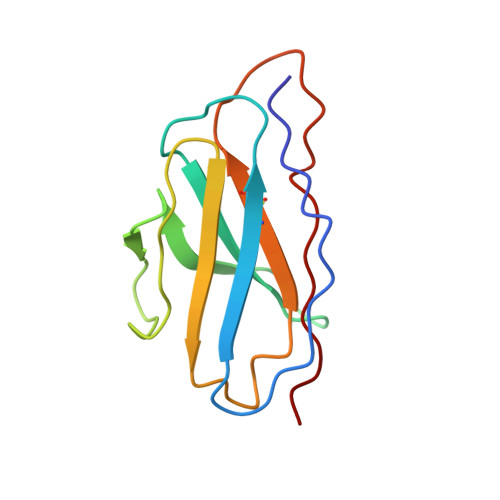
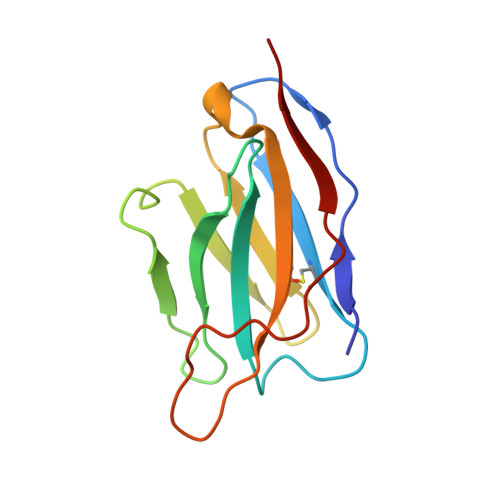
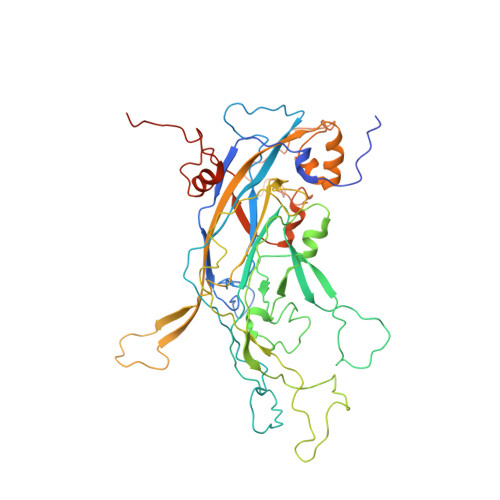
With more human papillomavirus (HPV) virus-like particle (VLP) vaccines to hit the market in future, a monoclonal antibody (mAb) with preferably comparable reactivity against vaccines from different expression systems and bioprocesses is urgently needed for the potency characterization. Among all mAbs against HPV16 collected, rabbit mAb H16.001 is potently neutralizing with the highest affinity, recognizes an immune-dominant epitope, and can comparably react with HPV16 vaccines from various sources. Cryo-electron microscopic (cryo-EM) structure demonstrated that 360 H16.001 Fabs could bind to HPV16 capsid in preferable binding manner without steric hindrance between neighboring Fabs, potentially supporting its identification for VLP structural integrity and utility in monitoring VLP structural probity. This structural analysis indicated that mAb H16.001 afforded unbiased potency characterization for various HPV16 vaccines and was potential for use in vaccine regulation practice. This study also showed a model process for selecting suitable mAbs for potency assays of other vaccines.
- Division of HIV/AIDS and Sexually Transmitted Virus Vaccines, National Institutes for Food and Drug Control (NIFDC), No.31 Huatuo Street, Daxing District, 102629 Beijing, China.
Organizational Affiliation: