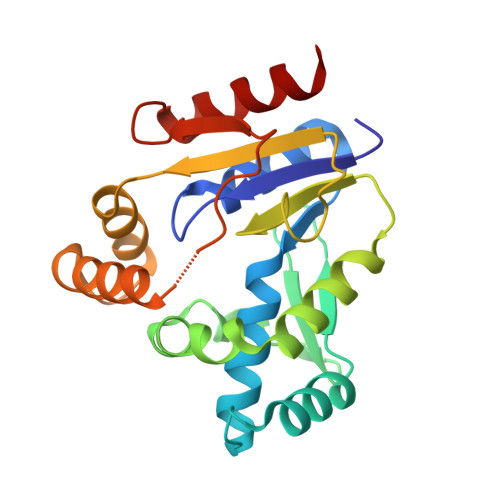
The crystal structures of Thermus thermophilus CMP kinase complexed with a phosphoryl group acceptor and donor.
Mega, R., Nakagawa, N., Kuramitsu, S., Masui, R.(2020) PLoS One 15: e0233689
- PubMed: 32469932 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0233689
- Primary Citation Related Structures:
7CKJ - PubMed Abstract:
Nucleoside monophosphate kinases play crucial roles in biosynthesis and regeneration of nucleotides. These are bi-substrate enzymes that catalyze reversible transfers of a phosphoryl group between ATP and nucleoside monophosphate. These enzymes are comprised of the CORE domain, the NMP-binding domain, and the LID domain. Large conformational rearrangement of the three domains occurs during the catalytic cycle. Although many structures of CMP kinase have been determined, only limited structural information has been available on the conformational changes along the reaction pathway. We determined five crystal structures of CMP kinase of Thermus thermophilus HB8 in ligand-free form and the CMP "open", CMP "closed", ADP-CDP-Gd3+-, and CDP-bound forms at resolutions of 1.7, 2.2, 1.5, 1.6, and 1.7 Å, respectively. The ligand-free form was in an open conformation, whereas the structures of the CMP "closed", ADP-CDP-Gd3+-, and CDP-bound forms were in a closed conformation, in which the shift of the NMP-binding domain and LID domain caused closure of the substrate-binding cleft. Interestingly, the CMP "open" form was in an open conformation even with CMP bound, implying intrinsic conformational fluctuation. The structure of the ADP-CDP complex is the first structure of CMP kinase with a phosphoryl group donor and an acceptor. Upon simultaneous binding of ADP and CDP, the side chains of several residues in the LID domain moved toward the nucleotides without global open-closed conformational changes compared to those in the CMP "closed" and CDP complexes. These global and local conformational changes may be crucial for the substrate recognition and catalysis. The terminal phosphate groups of ADP and CDP had similar geometry to those of two ADP in AMP kinase, suggesting common catalytic mechanisms to other nucleoside monophosphate kinases. Our findings are expected to contribute to detailed understanding of the reaction mechanism of CMP kinase.
- Graduate School of Frontier Biosciences, Osaka University, Suita, Osaka, Japan.
Organizational Affiliation: