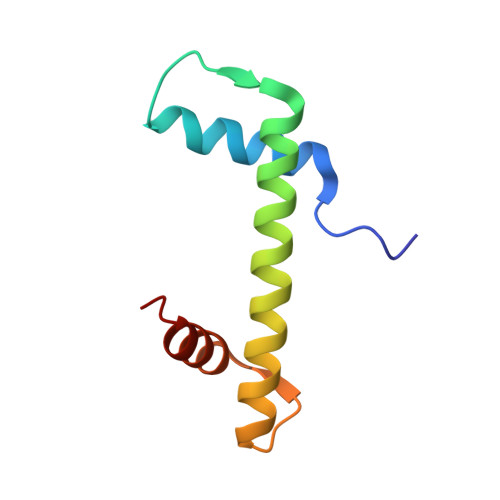
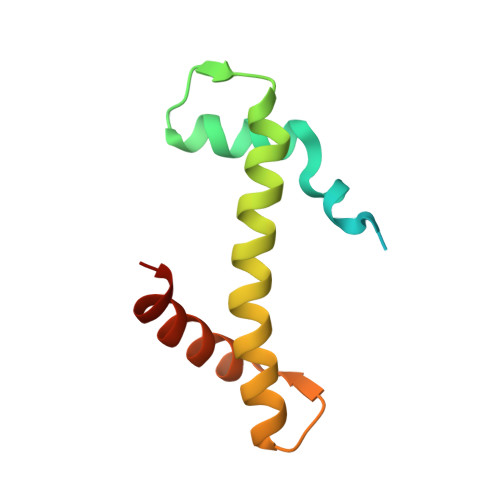
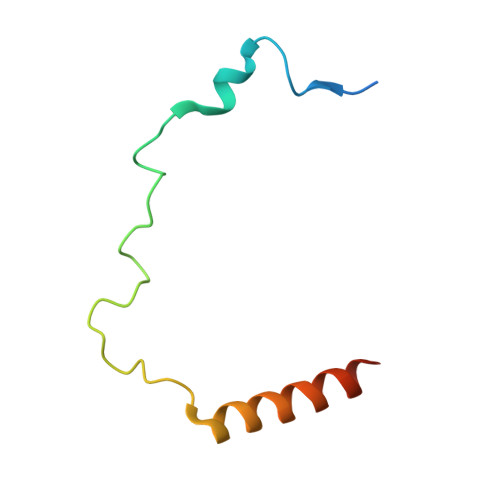
DNAJC9 integrates heat shock molecular chaperones into the histone chaperone network.
Hammond, C.M., Bao, H., Hendriks, I.A., Carraro, M., Garcia-Nieto, A., Liu, Y., Reveron-Gomez, N., Spanos, C., Chen, L., Rappsilber, J., Nielsen, M.L., Patel, D.J., Huang, H., Groth, A.(2021) Mol Cell 81: 2533-2548.e9
- PubMed: 33857403 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2021.03.041
- Primary Citation Related Structures:
7CIZ, 7CJ0 - PubMed Abstract:
From biosynthesis to assembly into nucleosomes, histones are handed through a cascade of histone chaperones, which shield histones from non-specific interactions. Whether mechanisms exist to safeguard the histone fold during histone chaperone handover events or to release trapped intermediates is unclear. Using structure-guided and functional proteomics, we identify and characterize a histone chaperone function of DNAJC9, a heat shock co-chaperone that promotes HSP70-mediated catalysis. We elucidate the structure of DNAJC9, in a histone H3-H4 co-chaperone complex with MCM2, revealing how this dual histone and heat shock co-chaperone binds histone substrates. We show that DNAJC9 recruits HSP70-type enzymes via its J domain to fold histone H3-H4 substrates: upstream in the histone supply chain, during replication- and transcription-coupled nucleosome assembly, and to clean up spurious interactions. With its dual functionality, DNAJC9 integrates ATP-resourced protein folding into the histone supply pathway to resolve aberrant intermediates throughout the dynamic lives of histones.
- Novo Nordisk Foundation Center for Protein Research (CPR), University of Copenhagen, Copenhagen, Denmark; Biotech Research and Innovation Centre (BRIC), University of Copenhagen, Copenhagen, Denmark.
Organizational Affiliation: