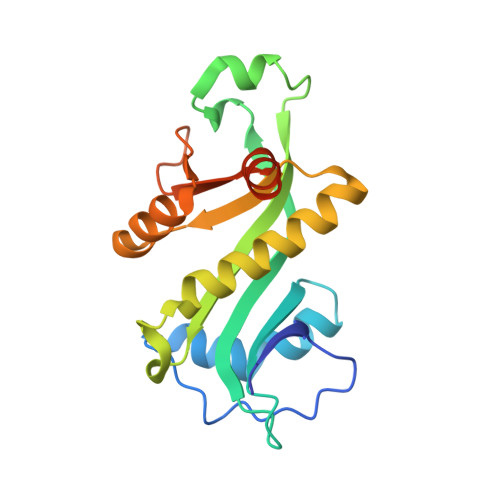
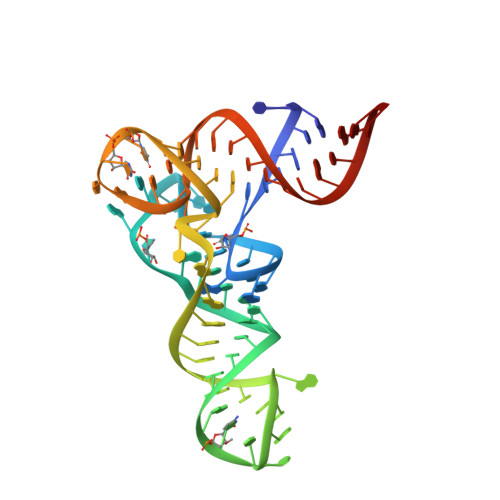
Mechanism of aminoacyl-tRNA acetylation by an aminoacyl-tRNA acetyltransferase AtaT from enterohemorrhagic E. coli.
Yashiro, Y., Sakaguchi, Y., Suzuki, T., Tomita, K.(2020) Nat Commun 11: 5438-5438
- PubMed: 33116145 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19281-z
- Primary Citation Related Structures:
7CHD - PubMed Abstract:
Toxin-antitoxin systems in bacteria contribute to stress adaptation, dormancy, and persistence. AtaT, a type-II toxin in enterohemorrhagic E. coli, reportedly acetylates the α-amino group of the aminoacyl-moiety of initiator Met-tRNAf Met , thus inhibiting translation initiation. Here, we show that AtaT has a broader specificity for aminoacyl-tRNAs than initially claimed. AtaT efficiently acetylates Gly-tRNA Gly , Trp-tRNA Trp , Tyr-tRNA Tyr and Phe-tRNA Phe isoacceptors, in addition to Met-tRNAf Met , and inhibits global translation. AtaT interacts with the acceptor stem of tRNAf Met , and the consecutive G-C pairs in the bottom-half of the acceptor stem are required for acetylation. Consistently, tRNA Gly , tRNA Trp , tRNA Tyr and tRNA Phe also possess consecutive G-C base-pairs in the bottom halves of their acceptor stems. Furthermore, misaminoacylated valyl-tRNAf Met and isoleucyl-tRNAf Met are not acetylated by AtaT. Therefore, the substrate selection by AtaT is governed by the specific acceptor stem sequence and the properties of the aminoacyl-moiety of aminoacyl-tRNAs.
- Department of Computational Biology and Medical Sciences, Graduate School of Frontier Sciences, The University of Tokyo, Kashiwa, Chiba, 277-8562, Japan.
Organizational Affiliation: