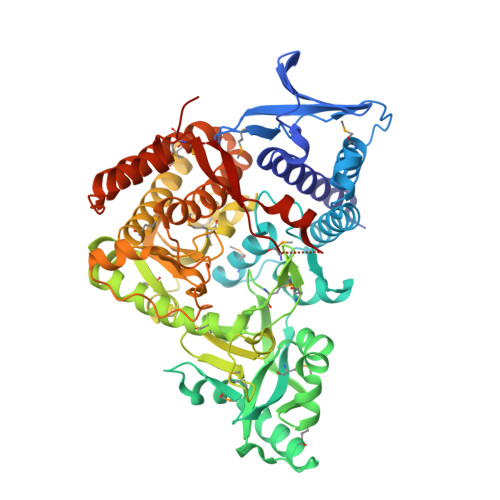
Structural and Biochemical Characterization of SbnC as a Representative Type B Siderophore Synthetase.
Tang, J., Ju, Y., Zhou, J., Guo, J., Gu, Q., Xu, J., Zhou, H.(2020) ACS Chem Biol 15: 2731-2740
- PubMed: 32880431 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.0c00523
- Primary Citation Related Structures:
7CBB - PubMed Abstract:
Staphyloferrin B is a key siderophore secreted by Staphylococcus aureus to acquire ferric ions from a host during infection, and its biosynthetic pathway has been validated to develop efficient antibacterial agents. Herein, we report the crystal structure of AMP-bound SbnC from S. aureus ( Sa SbnC) as the first representative structure of type B synthetases in the biosynthesis of α-hydroxycarboxylate siderophores. While type B synthetases specifically use α-ketoglutarate (α-KG) as their carboxylic acid substrate, Sa SbnC showed unique structural features in the substrate pocket compared with the type A and C synthetases. Screening of α-KG analogues suggested that the hydrogen-bonding interaction between the α-carbonyl group of α-KG and residue Lys552 is a key determinant for the substrate selectivity of type B synthetases. Interestingly, citrate, the product of the tricarboxylic acid cycle and the substrate of type A synthetases, was found to inhibit the activity of Sa SbnC with an IC 50 value of 83 μM by mimicking α-KG binding, suggesting a potential regulatory role of the tricarboxylic acid cycle, whose activity is under the control of the intracellular iron concentration, to Sa SbnC and other type B synthetases. These results provide critical new information to understand the structure, function, and regulation of type B synthetases in the siderophore-based iron acquisition system employed by a large number of pathogenic microbes.
- Research Center for Drug Discovery, School of Pharmaceutical Sciences, Sun Yat-sen University, Guangzhou 510006, China.
Organizational Affiliation: