Structural basis for transcription inhibition by E. coli SspA
Wei, L., Yu, F.(2020) Nucleic Acids Res
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Stringent starvation protein A | B [auth I], C [auth J] | 212 | Escherichia coli | Mutation(s): 0 Gene Names: sspA EC: 2.5.1.18 | 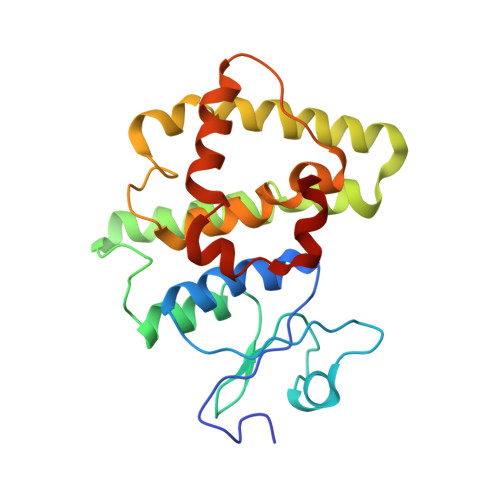 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0ACA3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA-directed RNA polymerase subunit alpha | D [auth K], E [auth A], F [auth B] | 329 | Escherichia coli | Mutation(s): 0 Gene Names: rpoA, ECPG_02615 EC: 2.7.7.6 | 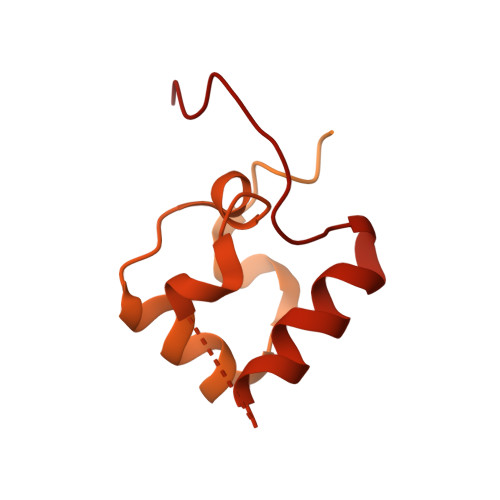 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A7Z4 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA-directed RNA polymerase subunit beta | G [auth C] | 1,342 | Escherichia coli K-12 | Mutation(s): 1 Gene Names: rpoB, groN, nitB, rif, ron, stl, stv, tabD, b3987, JW3950 EC: 2.7.7.6 | 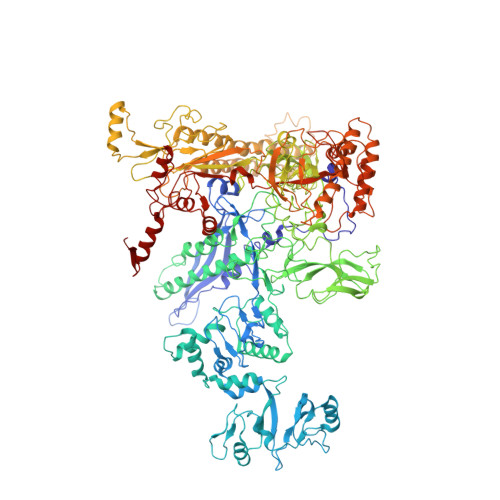 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A8V2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA-directed RNA polymerase subunit beta' | H [auth D] | 1,407 | Escherichia coli | Mutation(s): 0 Gene Names: rpoC EC: 2.7.7.6 | 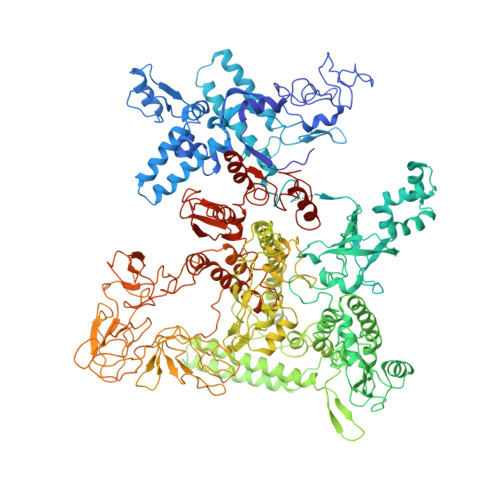 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A8T7 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 6 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA-directed RNA polymerase subunit omega | I [auth E] | 91 | Escherichia coli | Mutation(s): 0 Gene Names: rpoZ EC: 2.7.7.6 |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P0A800 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 7 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| RNA polymerase sigma factor RpoD | J [auth F] | 613 | Escherichia coli | Mutation(s): 0 Gene Names: rpoD |  |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00579 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 1 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (63-mer) | A [auth H] | 63 | Escherichia coli |  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
Entity ID: 8 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| DNA (63-mer) | K [auth G] | 63 | Escherichia coli |  |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ZN (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | M [auth D], N [auth D] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| MG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | L [auth D] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | RELION | 3.0 |