Target identification for small-molecule discovery in the FOXO3a tumor-suppressor pathway using a biodiverse peptide library.
Emery, A., Hardwick, B.S., Crooks, A.T., Milech, N., Watt, P.M., Mithra, C., Kumar, V., Giridharan, S., Sadasivam, G., Mathivanan, S., Sudhakar, S., Bairy, S., Bharatham, K., Hurakadli, M.A., Prasad, T.K., Kamariah, N., Muellner, M., Coelho, M., Torrance, C.J., McKenzie, G.J., Venkitaraman, A.R.(2021) Cell Chem Biol 28: 1602-1615.e9
- PubMed: 34111400 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2021.05.009
- Primary Citation Related Structures:
7C8E - PubMed Abstract:
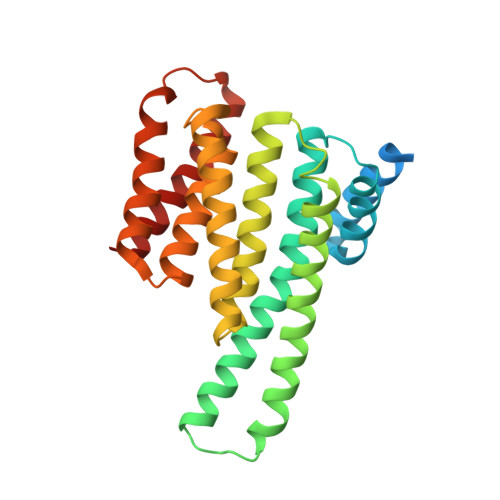
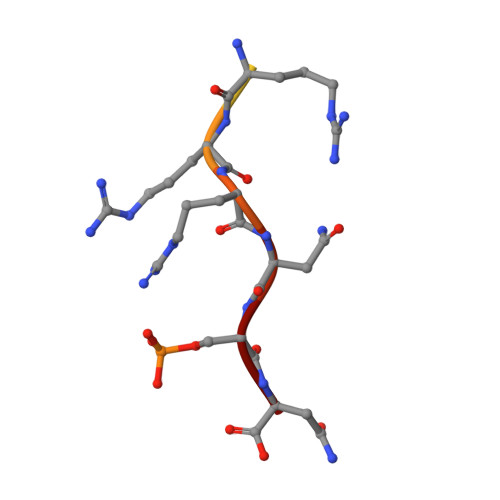
Genetic screening technologies to identify and validate macromolecular interactions (MMIs) essential for complex pathways remain an important unmet need for systems biology and therapeutics development. Here, we use a library of peptides from diverse prokaryal genomes to screen MMIs promoting the nuclear relocalization of Forkhead Box O3 (FOXO3a), a tumor suppressor more frequently inactivated by post-translational modification than mutation. A hit peptide engages the 14-3-3 family of signal regulators through a phosphorylation-dependent interaction, modulates FOXO3a-mediated transcription, and suppresses cancer cell growth. In a crystal structure, the hit peptide occupies the phosphopeptide-binding groove of 14-3-3ε in a conformation distinct from its natural peptide substrates. A biophysical screen identifies drug-like small molecules that displace the hit peptide from 14-3-3ε, providing starting points for structure-guided development. Our findings exemplify "protein interference," an approach using evolutionarily diverse, natural peptides to rapidly identify, validate, and develop chemical probes against MMIs essential for complex cellular phenotypes.
- Medical Research Council Cancer Unit, University of Cambridge, Hills Road, Cambridge CB2 0XZ, UK.
Organizational Affiliation: