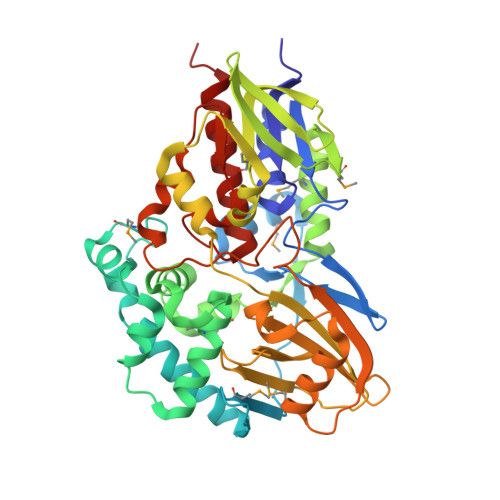
Molecular Deceleration Regulates Toxicant Release to Prevent Cell Damage in Pseudomonas putida S16 (DSM 28022).
Tang, H., Zhang, K., Hu, H., Wu, G., Wang, W., Zhu, X., Liu, G., Xu, P.(2020) mBio 11
- PubMed: 32873764 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.02012-20
- Primary Citation Related Structures:
7C49, 7C4A - PubMed Abstract:
The underlying molecular mechanisms of flavin-dependent amine oxidases remain relatively poorly understood, even though many of these enzymes have been reported. The nicotine oxidoreductase NicA2 is a crucial enzyme for the first step of nicotine degradation in Pseudomonas putida S16 (DSM 28022). Here, we present the crystal structure of a ternary complex comprising NicA2 residues 21 to 482, flavin adenine dinucleotide (FAD), and nicotine at 2.25 Å resolution. Unlike other, related structures, NicA2 does not have an associated diacyl glycerophospholipid, wraps its substrate more tightly, and has an intriguing exit passage in which nine bulky amino acid residues occlude the release of its toxic product, pseudooxynicotine (PN). The replacement of these bulky residues by amino acids with small side chains effectively increases the catalytic turnover rate of NicA2. Our results indicate that the passage in wild-type NicA2 effectively controls the rate of PN release and thus prevents its rapid intracellular accumulation. It gives ample time for PN to be converted to less-harmful substances by downstream enzymes such as pseudooxynicotine amine oxidase (Pnao) before its accumulation causes cell damage or even death. The temporal metabolic regulation mode revealed in this study may shed light on the production of cytotoxic compounds. IMPORTANCE Flavin-dependent amine oxidases have received extensive attention because of their importance in drug metabolism, Parkinson's disease, and neurotransmitter catabolism. However, the underlying molecular mechanisms remain relatively poorly understood. Here, combining the crystal structure of NicA2 (an enzyme in the first step of the bacterial nicotine degradation pathway in Pseudomonas putida S16 (DSM 28022)), biochemical analysis, and mutant construction, we found an intriguing exit passage in which bulky amino acid residues occlude the release of the toxic product of NicA2, in contrast to other, related structures. The selective product exportation register for NicA2 has proven to be beneficial to cell growth. Those seeking to produce cytotoxic compounds could greatly benefit from the use of such an export register mechanism.
- State Key Laboratory of Microbial Metabolism, Shanghai Jiao Tong University, Shanghai, People's Republic of China tanghongzhi@sjtu.edu.cn pingxu@sjtu.edu.cn.
Organizational Affiliation: