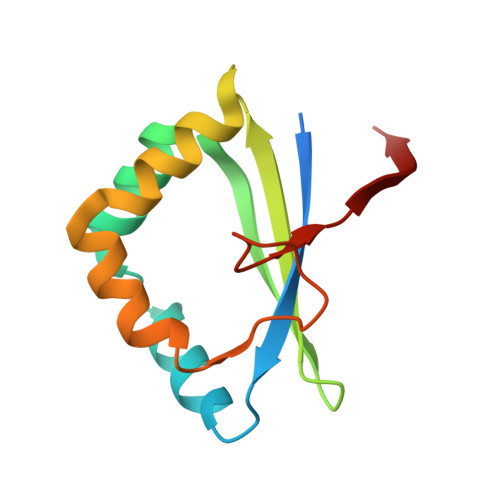
Functional and structural characterization of a novel L-fucose mutarotase involved in non-phosphorylative pathway of L-fucose metabolism.
Watanabe, Y., Watanabe, S., Fukui, Y., Nishiwaki, H.(2020) Biochem Biophys Res Commun 528: 21-27
- PubMed: 32448506 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.05.094
- Primary Citation Related Structures:
7BYU, 7BYW - PubMed Abstract:
Mutarotases catalyze the α-β anomeric conversion of monosaccharide, and play a key role in utilizing sugar as enzymes involved in sugar metabolism have specificity for the α- or β-anomer. In spite of the sequential similarity to l-rhamnose mutarotase protein superfamily (COG3254: RhaM), the ACAV_RS08160 gene in Acidovorax avenae ATCC 19860 (AaFucM) is located in a gene cluster related to non-phosphorylative l-fucose and l-galactose metabolism, and transcriptionally induced by these carbon sources; therefore, the physiological role remains unclear. Here, we report that AaFucM possesses mutarotation activity only toward l-fucose by saturation difference (SD) NMR experiments. Moreover, we determined the crystal structures of AaFucM in the apo form and in the l-fucose-bound form at resolutions of 2.21 and 1.75 Å, respectively. The overall structural folding was clearly similar to the RhaM members, differed from the known l-fucose mutarotase (COG4154: FucU), strongly indicating their convergent evolution. The structure-based mutational analyses suggest that Tyr18 is important for catalytic action, and that Gln87 and Trp99 are involved in the l-fucose-specific recognition.
- Department of Bioscience, Graduate School of Agriculture, Ehime University, 3-5-7 Tarumi, Matsuyama, Ehime, 790-8566, Japan; Faculty of Agriculture, Ehime University, 3-5-7 Tarumi, Matsuyama, Ehime, 790-8566, Japan.
Organizational Affiliation: