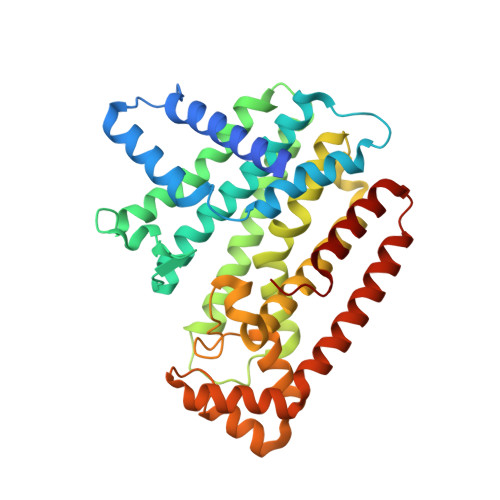
Structure-function studies of ultrahigh molecular weight isoprenes provide key insights into their biosynthesis.
Kajiura, H., Yoshizawa, T., Tokumoto, Y., Suzuki, N., Takeno, S., Takeno, K.J., Yamashita, T., Tanaka, S.I., Kaneko, Y., Fujiyama, K., Matsumura, H., Nakazawa, Y.(2021) Commun Biol 4: 215-215
- PubMed: 33594248 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-021-01739-5
- Primary Citation Related Structures:
7BUU, 7BUV, 7BUW, 7BUX - PubMed Abstract:
Some plant trans-1,4-prenyltransferases (TPTs) produce ultrahigh molecular weight trans-1,4-polyisoprene (TPI) with a molecular weight of over 1.0 million. Although plant-derived TPI has been utilized in various industries, its biosynthesis and physiological function(s) are unclear. Here, we identified three novel Eucommia ulmoides TPT isoforms-EuTPT1, 3, and 5, which synthesized TPI in vitro without other components. Crystal structure analysis of EuTPT3 revealed a dimeric architecture with a central hydrophobic tunnel. Mutation of Cys94 and Ala95 on the central hydrophobic tunnel no longer synthesizd TPI, indicating that Cys94 and Ala95 were essential for forming the dimeric architecture of ultralong-chain TPTs and TPI biosynthesis. A spatiotemporal analysis of the physiological function of TPI in E. ulmoides suggested that it is involved in seed development and maturation. Thus, our analysis provides functional and mechanistic insights into TPI biosynthesis and uncovers biological roles of TPI in plants.
- International Center for Biotechnology, Osaka University, 2-1 Yamada-oka, Suita, Osaka, 565-0871, Japan.
Organizational Affiliation: