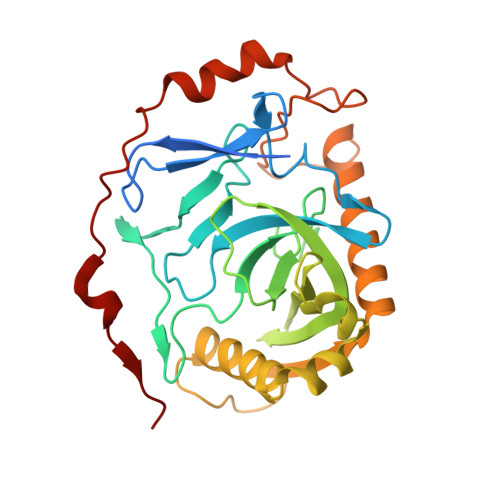
Structural and biochemical characterization of inorganic pyrophosphatase from Homo sapiens.
Hu, F., Huang, Z., Zheng, S., Wu, Q., Chen, Y., Lin, H., Huang, W., Li, L.(2020) Biochem Biophys Res Commun 533: 1115-1121
- PubMed: 33036755 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.09.139
- Primary Citation Related Structures:
7BTN, 7CMO - PubMed Abstract:
Inorganic pyrophosphatase (PPase) plays an essential role in energy conservation and provides energy for many biosynthetic pathways. Here, we present two three-dimensional structures of PPase from Homo sapiens (Hu-PPase) at 2.38 Å and 3.40 Å in different crystallization conditions. One of the Hu-PPase structures complex of two magnesium metal ions was determined to be a monomer (Hu-PPase-mono) here, while the other one to be a dimer-dimer (Hu-PPase-dd). In each asymmetric unit of Hu-PPase-mono, there are four α-helices and ten β-strands and folds as a barrel structure, and the active site contains two magnesium ions. Like PPases from many species, we found that Hu-PPase was able to undergo self-assembly. To our surprise, disruption of the self-assembly of Hu-PPase did not influence its enzymatic activity or the ability to promote cell growth. Our work uncovered that different structure forms of Hu-PPase and found that the pyrophosphatase activity of Hu-PPase is independent of its self-assembly.
- Key Laboratory of Gastrointestinal Cancer (Fujian Medical University), Ministry of Education, School of Basic Medical Sciences, Fujian Medical University, Fuzhou, China.
Organizational Affiliation: