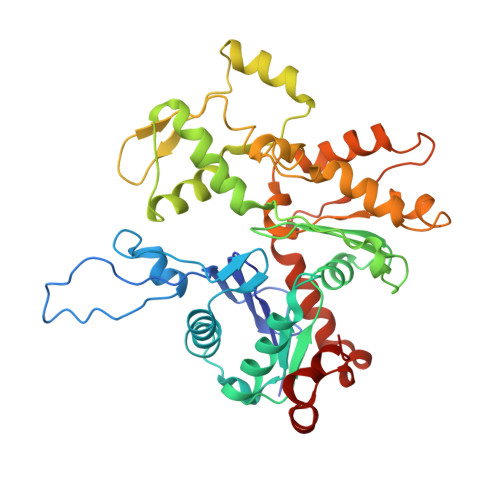
Structural insights into actin filament recognition by commonly used cellular actin markers.
Kumari, A., Kesarwani, S., Javoor, M.G., Vinothkumar, K.R., Sirajuddin, M.(2020) EMBO J 39: e104006-e104006
- PubMed: 32567727 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embj.2019104006
- Primary Citation Related Structures:
6M5G, 7BT7, 7BTE, 7BTI - PubMed Abstract:
Cellular studies of filamentous actin (F-actin) processes commonly utilize fluorescent versions of toxins, peptides, and proteins that bind actin. While the choice of these markers has been largely based on availability and ease, there is a severe dearth of structural data for an informed judgment in employing suitable F-actin markers for a particular requirement. Here, we describe the electron cryomicroscopy structures of phalloidin, lifeAct, and utrophin bound to F-actin, providing a comprehensive high-resolution structural comparison of widely used actin markers and their influence towards F-actin. Our results show that phalloidin binding does not induce specific conformational change and lifeAct specifically recognizes closed D-loop conformation, i.e., ADP-Pi or ADP states of F-actin. The structural models aided designing of minimal utrophin and a shorter lifeAct, which can be utilized as F-actin marker. Together, our study provides a structural perspective, where the binding sites of utrophin and lifeAct overlap with majority of actin-binding proteins and thus offering an invaluable resource for researchers in choosing appropriate actin markers and generating new marker variants.
- Center for Cardiovascular Biology and Diseases, Institute for Stem Cell Science and Regenerative Medicine, Bengaluru, India.
Organizational Affiliation: