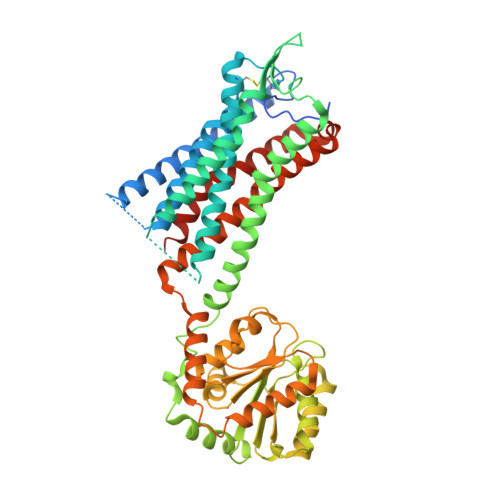
Structure of the human gonadotropin-releasing hormone receptor GnRH1R reveals an unusual ligand binding mode.
Yan, W., Cheng, L., Wang, W., Wu, C., Yang, X., Du, X., Ma, L., Qi, S., Wei, Y., Lu, Z., Yang, S., Shao, Z.(2020) Nat Commun 11: 5287-5287
- PubMed: 33082324 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19109-w
- Primary Citation Related Structures:
7BR3 - PubMed Abstract:
Gonadotrophin-releasing hormone (GnRH), also known as luteinizing hormone-releasing hormone, is the main regulator of the reproductive system, acting on gonadotropic cells by binding to the GnRH1 receptor (GnRH1R). The GnRH-GnRH1R system is a promising therapeutic target for maintaining reproductive function; to date, a number of ligands targeting GnRH1R for disease treatment are available on the market. Here, we report the crystal structure of GnRH1R bound to the small-molecule drug elagolix at 2.8 Å resolution. The structure reveals an interesting N-terminus that could co-occupy the enlarged orthosteric binding site together with elagolix. The unusual ligand binding mode was further investigated by structural analyses, functional assays and molecular docking studies. On the other hand, because of the unique characteristic of lacking a cytoplasmic C-terminal helix, GnRH1R exhibits different microswitch structural features from other class A GPCRs. In summary, this study provides insight into the ligand binding mode of GnRH1R and offers an atomic framework for rational drug design.
- Division of Nephrology and Kidney Research Institute, State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, Sichuan University, Chengdu, Sichuan, 610041, China.
Organizational Affiliation: