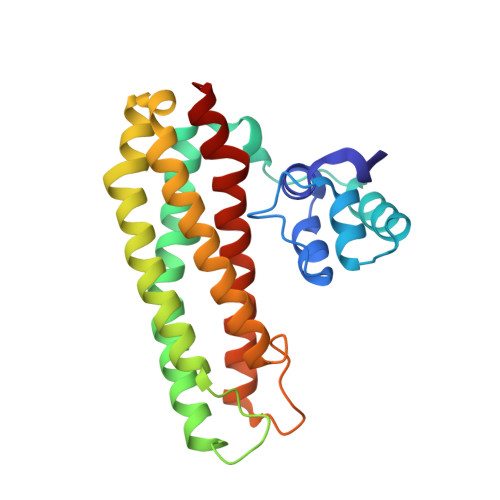
Structural Basis of RICs Iron Donation for Iron-Sulfur Cluster Biogenesis.
Silva, L.S.O., Matias, P.M., Romao, C.V., Saraiva, L.M.(2021) Front Microbiol 12: 670681-670681
- PubMed: 33995335 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmicb.2021.670681
- Primary Citation Related Structures:
7BHA, 7BHB, 7BHC - PubMed Abstract:
Escherichia coli YtfE is a di-iron protein of the widespread Repair of Iron Centers proteins (RIC) family that has the capacity to donate iron, which is a crucial component of the biogenesis of the ubiquitous family of iron-sulfur proteins. In this work we identify in E. coli a previously unrecognized link between the YtfE protein and the major bacterial system for iron-sulfur cluster (ISC) assembly. We show that YtfE establishes protein-protein interactions with the scaffold IscU, where the transient cluster is formed, and the cysteine desulfurase IscS. Moreover, we found that promotion by YtfE of the formation of an Fe-S cluster in IscU requires two glutamates, E125 and E159 in YtfE. Both glutamates form part of the entrance of a protein channel in YtfE that links the di-iron center to the surface. In particular, E125 is crucial for the exit of iron, as a single mutation to leucine closes the channel rendering YtfE inactive for the build-up of Fe-S clusters. Hence, we provide evidence for the key role of RICs as bacterial iron donor proteins involved in the biogenesis of Fe-S clusters.
- Instituto de Tecnologia Química e Biológica António Xavier, Universidade Nova de Lisboa, Oeiras, Portugal.
Organizational Affiliation: