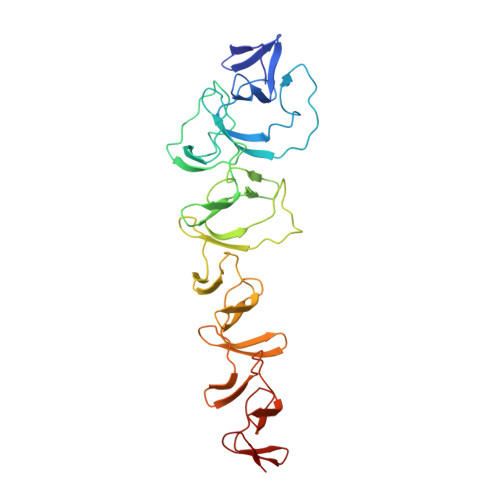
First Lanthanide Complex for De Novo Phasing in Native Protein Crystallography at 1 angstrom Radiation
Prieto-Castaneda, A., Martinez-Caballero, S., Agarrabeitia, A.R., Garcia-Moreno, I., Moya, S.D.L., Ortiz, M.J., Hermoso, J.A.(2021) ACS Appl Bio Mater 4: 4575-4581