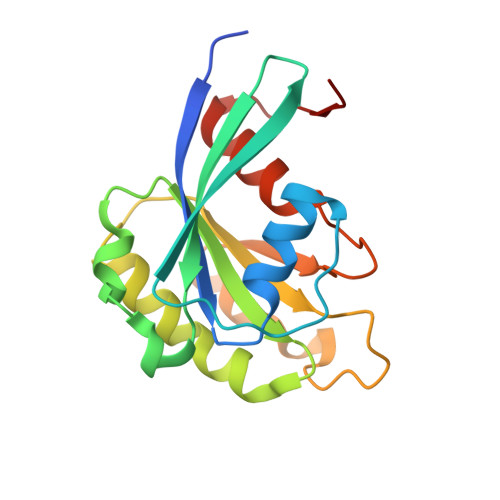
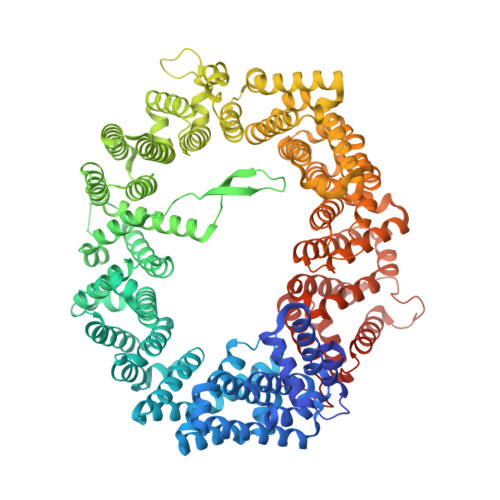
Crystal structure of human CRM1, covalently modified by 2-mercaptoethanol on Cys528, in complex with RanGTP.
Shaikhqasem, A., Schmitt, K., Valerius, O., Ficner, R.(2021) Acta Crystallogr F Struct Biol Commun 77: 70-78
- PubMed: 33682791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X2100203X
- Primary Citation Related Structures:
7B51 - PubMed Abstract:
CRM1 is a nuclear export receptor that has been intensively targeted over the last decade for the development of antitumor and antiviral drugs. Structural analysis of several inhibitor compounds bound to CRM1 revealed that their mechanism of action relies on the covalent modification of a critical cysteine residue (Cys528 in the human receptor) located in the nuclear export signal-binding cleft. This study presents the crystal structure of human CRM1, covalently modified by 2-mercaptoethanol on Cys528, in complex with RanGTP at 2.58 Å resolution. The results demonstrate that buffer components can interfere with the characterization of cysteine-dependent inhibitor compounds.
- Department for Molecular Structural Biology, Georg-August-Universität Göttingen, Justus-von-Liebig Weg 11, 37077 Göttingen, Germany.
Organizational Affiliation: