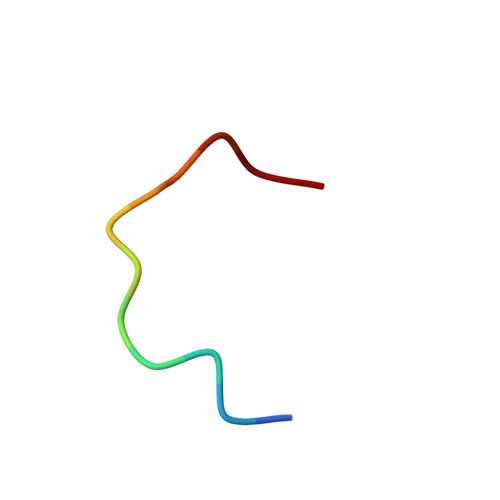
All - d - Enantiomeric Peptide D3 Designed for Alzheimer's Disease Treatment Dynamically Interacts with Membrane-Bound Amyloid-beta Precursors.
Bocharov, E.V., Gremer, L., Urban, A.S., Okhrimenko, I.S., Volynsky, P.E., Nadezhdin, K.D., Bocharova, O.V., Kornilov, D.A., Zagryadskaya, Y.A., Kamynina, A.V., Kuzmichev, P.K., Kutzsche, J., Bolakhrif, N., Muller-Schiffmann, A., Dencher, N.A., Arseniev, A.S., Efremov, R.G., Gordeliy, V.I., Willbold, D.(2021) J Med Chem 64: 16464-16479
- PubMed: 34739758 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c00632
- Primary Citation Related Structures:
7B3J, 7B3K - PubMed Abstract:
Alzheimer's disease (AD) is a severe neurodegenerative pathology with no effective treatment known. Toxic amyloid-β peptide (Aβ) oligomers play a crucial role in AD pathogenesis. All - d - Enantiomeric peptide D3 and its derivatives were developed to disassemble and destroy cytotoxic Aβ aggregates. One of the D3-like compounds is approaching phase II clinical trials; however, high-resolution details of its disease-preventing or pharmacological actions are not completely clear. We demonstrate that peptide D3 stabilizing Aβ monomer dynamically interacts with the extracellular juxtamembrane region of a membrane-bound fragment of an amyloid precursor protein containing the Aβ sequence. MD simulations based on NMR measurement results suggest that D3 targets the amyloidogenic region, not compromising its α-helicity and preventing intermolecular hydrogen bonding, thus creating prerequisites for inhibition of early steps of Aβ conversion into β-conformation and its toxic oligomerization. An enhanced understanding of the D3 action molecular mechanism facilitates development of effective AD treatment and prevention strategies.
- Research Center for Molecular Mechanisms of Aging and Age-related Diseases, Moscow Institute of Physics and Technology (State University), 141701 Dolgoprudny, Russia.
Organizational Affiliation: