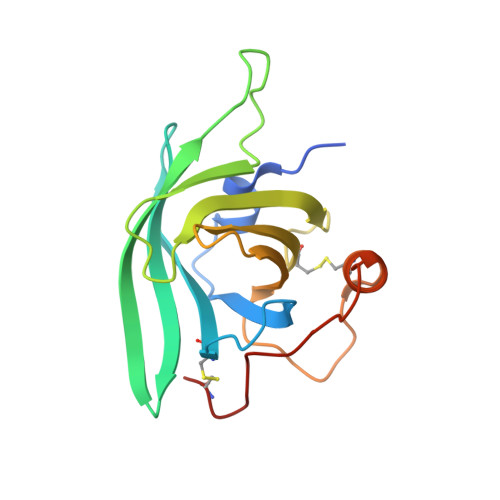
Structure and function of a family of tick-derived complement inhibitors targeting properdin.
Braunger, K., Ahn, J., Jore, M.M., Johnson, S., Tang, T.T.L., Pedersen, D.V., Andersen, G.R., Lea, S.M.(2022) Nat Commun 13: 317-317
- PubMed: 35031611 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-27920-2
- Primary Citation Related Structures:
7B26, 7B28, 7B29, 7B2A, 7B2D - PubMed Abstract:
Activation of the serum-resident complement system begins a cascade that leads to activation of membrane-resident complement receptors on immune cells, thus coordinating serum and cellular immune responses. Whilst many molecules act to control inappropriate activation, Properdin is the only known positive regulator of the human complement system. By stabilising the alternative pathway C3 convertase it promotes complement self-amplification and persistent activation boosting the magnitude of the serum complement response by all triggers. In this work, we identify a family of tick-derived alternative pathway complement inhibitors, hereafter termed CirpA. Functional and structural characterisation reveals that members of the CirpA family directly bind to properdin, inhibiting its ability to promote complement activation, and leading to potent inhibition of the complement response in a species specific manner. We provide a full functional and structural characterisation of a properdin inhibitor, opening avenues for future therapeutic approaches.
- Sir William Dunn School of Pathology, University of Oxford, OX1 3RE, Oxford, UK.
Organizational Affiliation: