Molecular rationale for antibody-mediated targeting of the hantavirus fusion glycoprotein.
Rissanen, I., Stass, R., Krumm, S.A., Seow, J., Hulswit, R.J., Paesen, G.C., Hepojoki, J., Vapalahti, O., Lundkvist, A., Reynard, O., Volchkov, V., Doores, K.J., Huiskonen, J.T., Bowden, T.A.(2020) Elife 9
- PubMed: 33349334 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.58242
- Primary Citation Related Structures:
6Z06, 7B09, 7B0A - PubMed Abstract:
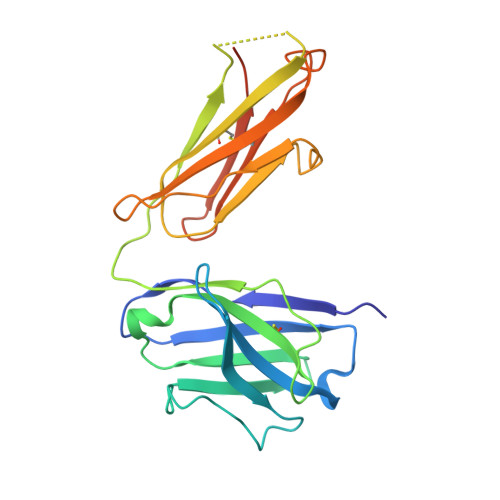
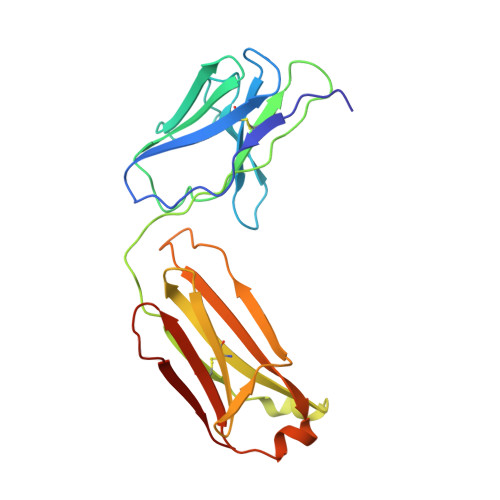
The intricate lattice of Gn and Gc glycoprotein spike complexes on the hantavirus envelope facilitates host-cell entry and is the primary target of the neutralizing antibody-mediated immune response. Through study of a neutralizing monoclonal antibody termed mAb P-4G2, which neutralizes the zoonotic pathogen Puumala virus (PUUV), we provide a molecular-level basis for antibody-mediated targeting of the hantaviral glycoprotein lattice. Crystallographic analysis demonstrates that P-4G2 binds to a multi-domain site on PUUV Gc and may preclude fusogenic rearrangements of the glycoprotein that are required for host-cell entry. Furthermore, cryo-electron microscopy of PUUV-like particles in the presence of P-4G2 reveals a lattice-independent configuration of the Gc, demonstrating that P-4G2 perturbs the (Gn-Gc) 4 lattice. This work provides a structure-based blueprint for rationalizing antibody-mediated targeting of hantaviruses.
- Division of Structural Biology, Wellcome Centre for Human Genetics, University of Oxford, Oxford, United Kingdom.
Organizational Affiliation: