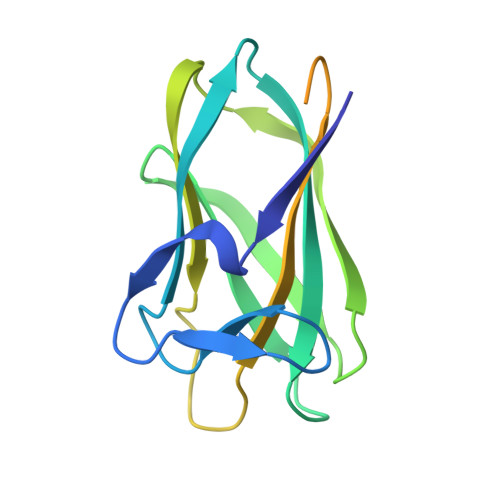
Structural and biophysical characterization of the multidomain xylanase Xyl.
Anye, V., Kruger, R.F., Schubert, W.D.(2022) PLoS One 17: e0269188-e0269188
- PubMed: 35657930 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0269188
- Primary Citation Related Structures:
7AX7, 7AY3, 7AYP, 7ZSZ - PubMed Abstract:
The depletion of fossil fuels, associated pollution, and resulting health hazards are of concern worldwide. Woody biomass constitutes an alternative source of cleaner and renewable energy. The efficient use of woody biomass depends on xylan depolymerisation as the endo-β-1,4-xylopyranosyl homopolymer is the main component of hemicellulose, the second most abundant component of wood. Xylan depolymerisation is achieved by hemicellulolytic xylanases of glycoside hydrolase (GH) families 5, 8, 10, 11, 30 and 43 of the CAZY database. We analysed a multidomain xylanase (Xyl) from the hindgut metagenome of the snouted harvester termite Trinervitermes trinervoides that releases xylobiose and xylotriose from beech and birch xylan and wheat arabinoxylan. The four domains of Xyl include an N-terminal GH11 xylanase domain, two family 36-like carbohydrate-binding domains CBM36-1 and 2, and a C-terminal CE4 esterase domain. Previous analyses indicated that CBM36-1 deletion slightly increased GH11 catalysis at low pH whereas removal of both CBMs decreased xylanase activity at 60°C from 90 to 56%. Possible cooperativity between the domains suggested by these observations was explored. A crystal structure of the two-domain construct, GH11-CBM36-1, confirmed the structure of the GH11 domain whereas the CBM36-1 domain lacked electron density, possibly indicating a random orientation of the CBM36-1 domain around the GH11 domain. Isothermal titration calorimetry (ITC) experiments similarly did not indicate specific interactions between the individual domains of Xyl supporting a "beads-on-a-string" model for Xyl domains.
- Department of Biochemistry, Genetics and Microbiology, Faculty of Natural and Agricultural Sciences, University of Pretoria, Pretoria, South Africa.
Organizational Affiliation: