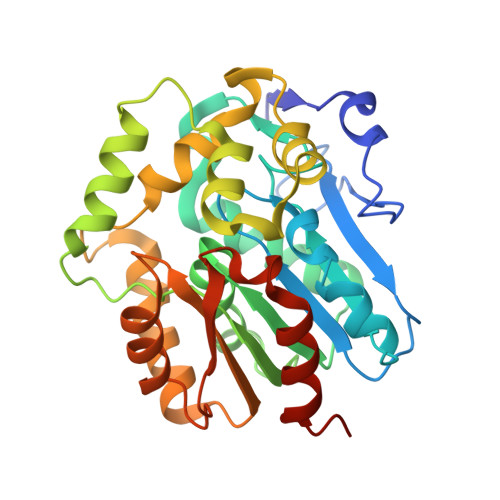
The tetrameric structure of the novel haloalkane dehalogenase DpaA from Paraglaciecola agarilytica NO2.
Mazur, A., Prudnikova, T., Grinkevich, P., Mesters, J.R., Mrazova, D., Chaloupkova, R., Damborsky, J., Kuty, M., Kolenko, P., Kuta Smatanova, I.(2021) Acta Crystallogr D Struct Biol 77: 347-356
- PubMed: 33645538 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798321000486
- Primary Citation Related Structures:
7AVR - PubMed Abstract:
Haloalkane dehalogenases (EC 3.8.1.5) are microbial enzymes that catalyse the hydrolytic conversion of halogenated compounds, resulting in a halide ion, a proton and an alcohol. These enzymes are used in industrial biocatalysis, bioremediation and biosensing of environmental pollutants or for molecular tagging in cell biology. The novel haloalkane dehalogenase DpaA described here was isolated from the psychrophilic and halophilic bacterium Paraglaciecola agarilytica NO2, which was found in marine sediment collected from the East Sea near Korea. Gel-filtration experiments and size-exclusion chromatography provided information about the dimeric composition of the enzyme in solution. The DpaA enzyme was crystallized using the sitting-drop vapour-diffusion method, yielding rod-like crystals that diffracted X-rays to 2.0 Å resolution. Diffraction data analysis revealed a case of merohedral twinning, and subsequent structure modelling and refinement resulted in a tetrameric model of DpaA, highlighting an uncommon multimeric nature for a protein belonging to haloalkane dehalogenase subfamily I.
- Faculty of Science, University of South Bohemia in Ceske Budejovice, Branisovska 1760, 370 05 Ceske Budejovice, Czech Republic.
Organizational Affiliation: