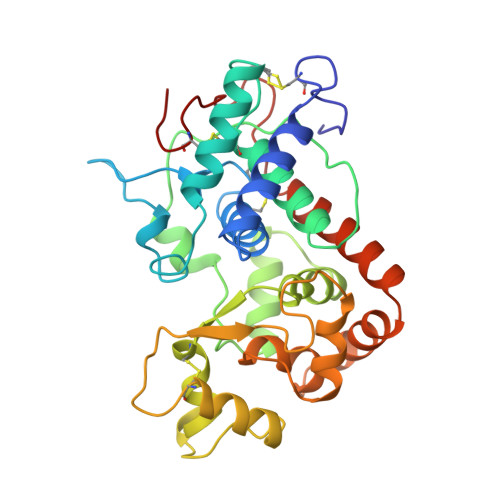
The structures of the horseradish peroxidase C-ferulic acid complex and the ternary complex with cyanide suggest how peroxidases oxidize small phenolic substrates.
Henriksen, A., Smith, A.T., Gajhede, M.(1999) J Biological Chem 274: 35005-35011
- PubMed: 10574977 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.49.35005
- Primary Citation Related Structures:
6ATJ, 7ATJ - PubMed Abstract:
We have solved the x-ray structures of the binary horseradish peroxidase C-ferulic acid complex and the ternary horseradish peroxidase C-cyanide-ferulic acid complex to 2.0 and 1.45 A, respectively. Ferulic acid is a naturally occurring phenolic compound found in the plant cell wall and is an in vivo substrate for plant peroxidases. The x-ray structures demonstrate the flexibility and dynamic character of the aromatic donor binding site in horseradish peroxidase and emphasize the role of the distal arginine (Arg(38)) in both substrate oxidation and ligand binding. Arg(38) hydrogen bonds to bound cyanide, thereby contributing to the stabilization of the horseradish peroxidase-cyanide complex and suggesting that the distal arginine will be able to contribute with a similar interaction during stabilization of a bound peroxy transition state and subsequent O-O bond cleavage. The catalytic arginine is additionally engaged in an extensive hydrogen bonding network, which also includes the catalytic distal histidine, a water molecule and Pro(139), a proline residue conserved within the plant peroxidase superfamily. Based on the observed hydrogen bonding network and previous spectroscopic and kinetic work, a general mechanism of peroxidase substrate oxidation is proposed.
- Protein Structure Group, Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Kobenhavn O, Denmark. anette@jerne.ki.ku.dk
Organizational Affiliation: