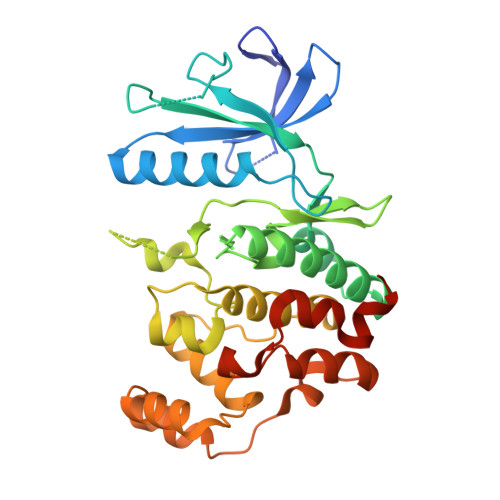
Crystal Structure and Inhibitor Identifications Reveal Targeting Opportunity for the Atypical MAPK Kinase ERK3.
Schroder, M., Filippakopoulos, P., Schwalm, M.P., Ferrer, C.A., Drewry, D.H., Knapp, S., Chaikuad, A.(2020) Int J Mol Sci 21
- PubMed: 33114754 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/ijms21217953
- Primary Citation Related Structures:
7AK3, 7AQB - PubMed Abstract:
Extracellular signal-regulated kinase 3 (ERK3), known also as mitogen-activated protein kinase 6 (MAPK6), is an atypical member of MAPK kinase family, which has been poorly studied. Little is known regarding its function in biological processes, yet this atypical kinase has been suggested to play important roles in the migration and invasiveness of certain cancers. The lack of tools, such as a selective inhibitor, hampers the study of ERK3 biology. Here, we report the crystal structure of the kinase domain of this atypical MAPK kinase, providing molecular insights into its distinct ATP binding pocket compared to the classical MAPK ERK2, explaining differences in their inhibitor binding properties. Medium-scale small molecule screening identified a number of inhibitors, several of which unexpectedly exhibited remarkably high inhibitory potencies. The crystal structure of CLK1 in complex with CAF052, one of the most potent inhibitors identified for ERK3, revealed typical type-I binding mode of the inhibitor, which by structural comparison could likely be maintained in ERK3. Together with the presented structural insights, these diverse chemical scaffolds displaying both reversible and irreversible modes of action, will serve as a starting point for the development of selective inhibitors for ERK3, which will be beneficial for elucidating the important functions of this understudied kinase.
- Structural Genomics Consortium, Goethe University Frankfurt, Buchmann Institute for Molecular Life Sciences, Max-von-Laue-Straße 15, 60438 Frankfurt am Main, Germany.
Organizational Affiliation: