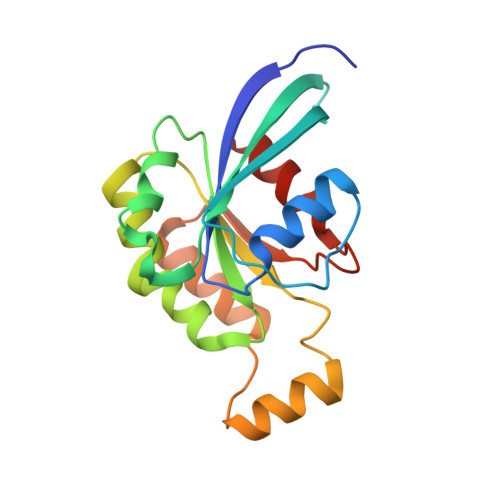
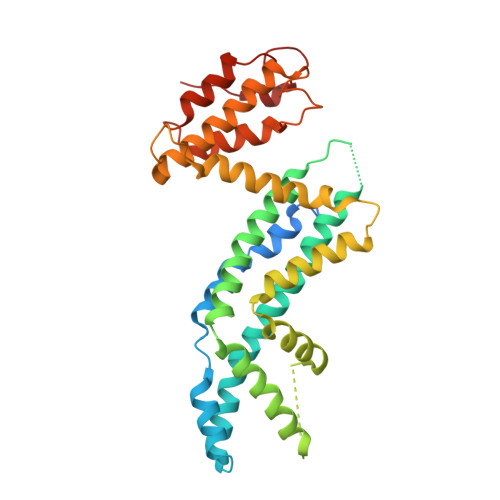
Structural Basis of CYRI-B Direct Competition with Scar/WAVE Complex for Rac1.
Yelland, T., Le, A.H., Nikolaou, S., Insall, R., Machesky, L., Ismail, S.(2021) Structure 29: 226-237.e4
- PubMed: 33217330 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2020.11.003
- Primary Citation Related Structures:
7AJK, 7AJL - PubMed Abstract:
Rac1 is a major regulator of actin dynamics, with GTP-bound Rac1 promoting actin assembly via the Scar/WAVE complex. CYRI competes with Scar/WAVE for interaction with Rac1 in a feedback loop regulating actin dynamics. Here, we reveal the nature of the CYRI-Rac1 interaction, through crystal structures of CYRI-B lacking the N-terminal helix (CYRI-BΔN) and the CYRI-BΔN:Rac1Q61L complex, providing the molecular basis for CYRI-B regulation of the Scar/WAVE complex. We reveal CYRI-B as having two subdomains - an N-terminal Rac1 binding subdomain with a unique Rac1-effector interface and a C-terminal Ratchet subdomain that undergoes conformational changes induced by Rac1 binding. Finally, we show that the CYRI protein family, CYRI-A and CYRI-B can produce an autoinhibited hetero- or homodimers, adding an additional layer of regulation to Rac1 signaling.
- CRUK- Beatson Institute, Glasgow G61 1BD, UK.
Organizational Affiliation: