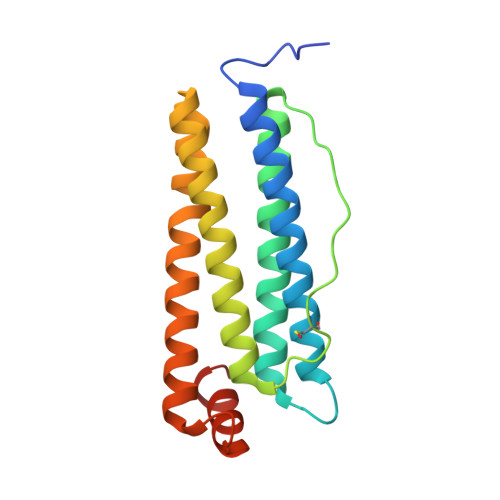
Atomic-resolution protein structure determination by cryo-EM.
Yip, K.M., Fischer, N., Paknia, E., Chari, A., Stark, H.(2020) Nature 587: 157-161
- PubMed: 33087927 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-2833-4
- Primary Citation Related Structures:
6Z6U, 6Z9E, 6Z9F, 7A6A, 7A6B - PubMed Abstract:
Single-particle electron cryo-microscopy (cryo-EM) is a powerful method for solving the three-dimensional structures of biological macromolecules. The technological development of transmission electron microscopes, detectors and automated procedures in combination with user-friendly image processing software and ever-increasing computational power have made cryo-EM a successful and expanding technology over the past decade 1 . At resolutions better than 4 Å, atomic model building starts to become possible, but the direct visualization of true atomic positions in protein structure determination requires much higher (better than 1.5 Å) resolution, which so far has not been attained by cryo-EM. The direct visualization of atom positions is essential for understanding the mechanisms of protein-catalysed chemical reactions, and for studying how drugs bind to and interfere with the function of proteins 2 . Here we report a 1.25 Å-resolution structure of apoferritin obtained by cryo-EM with a newly developed electron microscope that provides, to our knowledge, unprecedented structural detail. Our apoferritin structure has almost twice the 3D information content of the current world record reconstruction (at 1.54 Å resolution 3 ). We can visualize individual atoms in a protein, see density for hydrogen atoms and image single-atom chemical modifications. Beyond the nominal improvement in resolution, we also achieve a substantial improvement in the quality of the cryo-EM density map, which is highly relevant for using cryo-EM in structure-based drug design.
- Department of Structural Dynamics, MPI for Biophysical Chemistry, Göttingen, Germany.
Organizational Affiliation: